Dave Gray's Build-A-Gene Class Notes - Session 4
In session 4 we did ligation of our vectors and transformation of E. coli. We also had a discussion of current work in molecular biology.
Ligation
At the end of the previous session, we added restriction enzymes to our vector, promoter and emGFP gene and left them to work so that the restriction enzymes could snip the ends of each of these, leaving "sticky ends" that would allow the three pieces to be joined into a continuous loop, initially by hydrogen bonds due to the attraction of the nucleotides, then with covalent bonds by a process called ligation that "zips up" the DNA backbone at the junction points.
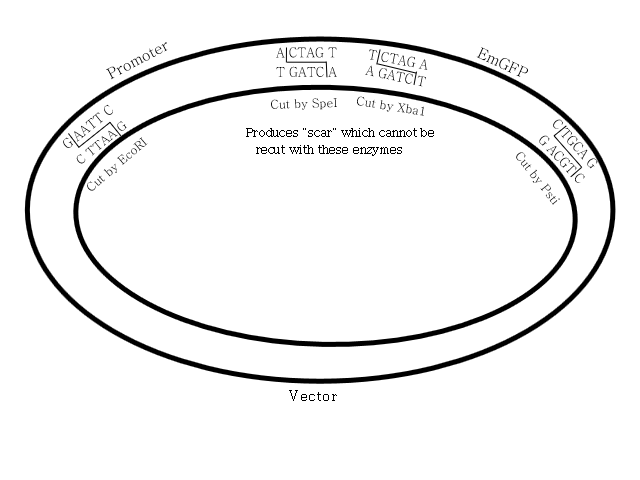
To perform this, we combined all three of our components with T4 ligase + buffer solution and incubated, first for 30 minutes at 16°C, then for 20 minutes at 80°C.
Lisa explained the rationale for the restriction enzymes that were chosen and some options for making this all work.
Some of the "tweaking" can be done in a DNA sequence to facilitate the use of restriction enzymes. For example, when planning this, you would have to ensure that there were no other occurrences of the nucleotide sequence that a restriction enzyme matches to such that it would cut your component where you don't want. The way nucleotides are matched to amino acids in assembling proteins offers one solution. In coding for amino acids, a set of three nucleotides is always selects a specific amino acid. However, many amino acids can be selected by more than one nucleotide sequence. So one triplet of nucleotides could be replaced with another that codes for the same amino acid without breaking the function of the genome.
This also explains one possible reason why we used three components. Logically, we could have combined the vector and promoter into a single strand. However, if PstI could cut the promoter at a location we don't want or SpeI would cut the vector at a location we didn't want, that would be a problem. However, by handling them as separate strands, that possibility is avoided. Before combining components, we heat them separately to 80°C. This deactivates the enzymes so that the components can be safely mixed.
We talked a bit about how restriction enzymes work. One point was that in the double helix of DNA, there is also a second double helix of "grooves" in the space between the backbones of the two strands. One side is larger and the other is smaller. These are referred to as "major" and "minor" grooves. Restriction enzymes tend to bind at points in the major groove. (It allows more space for them to access the nucleotides.)
Lisa explained that a vector like the one we constructed could also be used in yeast or mammalian cells. For it to be used there, we would have to add an appropriate origin of replication (origin or ori) for one or both of these.
It turns out that only a very small number of bacteria take up our vector. (e.g. 1/1,000,000) For that reason, we need a LOT of bacteria and a LOT of vectors. To weed out those that have from those that have not, our vector includes a selectable marker. We used the Chlorr gene. It allows the bacteria to live in the presence of the antibiotic chloramphenicol. Then, by including that antibiotic in the growth material in the petri dishes, we ensure that only bacteria with our vector will survive and grow. (Many of these will have flaws in the vector and so will not fluoresce green. However, they will be far more concentrated.
A tip from the class - always label the bottom of the petri dish, not the top. That way, if you drop them and the lid goes rolling off, you haven't lost your label.
partsregistry.org
Lisa mentioned the standard registry of biological parts at partsregistry.org (now http://parts.igem.org/Main_Page). This allows for labs to share reusable parts. (Not all work as advertised.) She plans to submit our EmGFP gene to the registry after our class is complete.
IGEM - International Genetically Engineered Competition
We discussed the IGEM competition, a competition for genetically engineered projects. Two commercial spinoffs from IGEM are Ginko Bioworks and BioFab.
Synthetic Bacteria
During this session, Lisa discussed the work done by Craig Venter to create bacteria with a synthetic genome. A well known saying from Venter is that the genome is the only software that produces its own hardware.
Reasons for doing this:
- Minimal Genome - This allows us to determine what genes are actually needed for a free living organism to function.
- By eliminating non-essential functions, can maximize output of desired product vs. other cell activities.
In 2010 Venter's institute announced they had successfully produced a working synthetic genome for bacteria. It their work, they started trying to produce a synthetic genome for m. genitalium. This is the smallest free living bacteria. To test whether the synthetic genome would function, they added it to m. capricolum, another bacteria, to see whether the genome would stimulate the production of m. genitalium. This effort lasted 10 years and was unsuccessful.
Next, they tried m. mycoides. This is the next smallest and has the benefit of growing faster. For two years, this also didn't work. Then they realized the problem was that the restriction enzymes were breaking down their genome. By adding methyl groups to their DNA, they were able to protect it from the restriction enzymes and, as a result, succeeded. (As a follow-up, I asked why we didn't run into the same issue in our effort to insert DNA into bacteria. Lisa explained that the lab strain of E. coli we are working with have been genetically modified to disable the restriction enzymes. Venter's team did not do so.)
Their process used a technique called "Gibson Assembly" for assembling the genome. They start with strands with only about 1000 base pairs. They add exonuclease 1, polymerase and ligase. This cuts one strand and nibbles back the nucleotides. Then strands can assemble and polymerize the gaps and ligate.
Then they take 10 fragments and a yeast vector. The yeast uses homologous recombination to attach the fragments - a process that is more complex than we were ready to discuss. Homologous recombination is a normal part of DNA repair.