Dave Gray's Build-A-Gene Class Notes - Session 2
In session 2, we assembled the emGFP coding sequence from smaller segments and amplified the result. We also used gel electrophoresis to check the results of our Session 1 PCR.
Polymerase Cycling Assembly
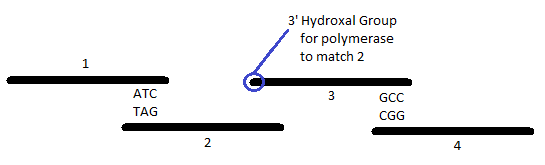
The emGFP coding sequence is about 750 nucleotides long. We ordered this in 60 nucleotide sets - about 20 pieces. That means that what we ordered was about 1200 nucleotides. The reason this is greater than the 750 nucleotide target is that the pieces are designed to overlap by about 20 nucleotides so that the segments will line up correctly. Each strand includes a 3' OH for the polymerase to attach to so that it can fill in the gaps. We use PCA (Polymerase Cycling Assembly) to accomplish this. Controlling the temperature helps ensure that segments with flaws in the overlapping sections do not successfully anneal. (Tm = melting temperature = temperature at which 50% of segments anneal and 50% fail. We set an annealing temp at Tm - 5°C.) In the end, we will have a little of what we want and a lot of what we don't want - sequences with flaws.
The reason for ordering shorter segments is that it helps ensure that we get some error free emGFP coding sequences. The error rate for chemically synthesizing DNA is .995 (5/1000). So in a 1000 nucleotide segment, on average, we are likely to get 5 errors. Or for 750 nucleotides (for emGFP), the number would be closer to 4.
Another way of calculating this is use the probability of a correct outcome for each nucleotide added - .995 or 99.5%. To calculate the likelihood of any series of nucleotides being correct, we need to raise .995 to the power of the number of nucleotides. So for a 750 nucleotide emGFP sequence, we would calculate (.995)750 = .027 or 2.7% of the resulting strands being correct. By reducing the length to 60, we get (.995)60 = .74 or 74% of the segments are good. We will amplify these to produce many copies and then join them. As a result, we should have many perfect sequences - along with many more imperfect. So we will have to weed out the ones we want.
Chemical synthesis of DNA uses phosphoramidite nucleotides, a modified precursor to DNA. Later we end up with real DNA.
We will be using herculase rather than taq polymerase because of it's better error rate. (My notes say 1 in 8 vs 1 in 12, but unclear what this means.)
Amplifying the emGFP
After using PCA to assemble the fragments, we used PCR to amplify the full-length strands. This weeds out shorter strands since, unless they belong to an end of the emGFP, the primers won't attach to them and unless they have a primer on both end, they won't amplify.
Designing the segments
Determining how to break up a desired DNA segment into shorter pieces with overlapping sections can be tricky, but there is free software to help with this. We looked at a program call Gene Design (found at genedesign.org). We used the feature, "Building Block Design."
Gel Electrophoresis
Gel Electrophoresis is a technique for sorting strands of DNA by their length in a way that can then be evaluated visually. The technique begins with placing the DNA strands mixed with a blue "loading dye" in a small "well" (hole) in one end of a gel that has been submerged TAE (Tris-acetate-EDTA) buffer. The loading dye makes it easier to see whether you have place the DNA correctly and helps weigh the DNA down so that it says in the well rather than flowing out into the TAE buffer. The well is positioned near the negative connection of an electrode. When the electricity is turned on, the DNA is attracted to the positive charge at the other end of the gel. (The DNA backbone, made up of phosphate groups, is located to the outside of the double helix, giving it a negative charge.) The gel is a sort of carbohydrate mesh made from seaweed. Because smaller strands of DNA can find their way through the gel more quickly, after some period of time, the smaller strands will be much closer to the positive terminal than longer strands.
Alongside the sample being tested, a "ladder", a known mix of DNA molecules is placed in a separate well. The "ladder" will form visible reference points in the gel showing where strands of known length have migrated to. The gel includes GelRed which merges into the double helix between the bases and fluoresces red under UV light making the results easy to see. Since our vector should be 2070 base pairs and our promoter + RBS should be 500 base pairs, we were able to see whether the results confirmed that our first round of PCR produced the expected result.