User:Trisha I. Ibeh/Notebook/Trisha Notebook/2013/09/03: Difference between revisions
(→Notes) |
|||
| (16 intermediate revisions by the same user not shown) | |||
| Line 49: | Line 49: | ||
The RANDOM concentration was 0.25*10^-5 for adenosine and 0.4*10^-5 for inosine. | The RANDOM concentration was 0.25*10^-5 for adenosine and 0.4*10^-5 for inosine. | ||
'''Preparation of Dilutions''' | '''Preparation of Dilutions of Adenosine''' | ||
[[Image:Screen_Shot_2013-10-05_at_4.20.59_AM.png]]<br> | |||
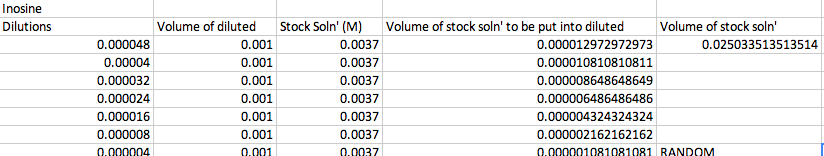
'''Preparation of Dilutions of Inosine''' | |||
[[Image:Screen_Shot_2013-10-05_at_3.41.48_AM.png]]<br> | |||
The groups then exchanged unknowns to determine the concentration from the calibration curves. In a week, the data will be revisited and the error will be propagated from the calibration curve to the concentration calculation. After making the calculation, the calculation of the unknown from the group will be compared. | |||
[[Category:Course]] | [[Category:Course]] | ||
| Line 66: | Line 65: | ||
[[Category:Course]] | [[Category:Course]] | ||
[[Category:Miscellaneous]] | [[Category:Miscellaneous]] | ||
==Data== | ==Data== | ||
'''Adenosine Absorbance Spectrum''' | '''Adenosine Absorbance Spectrum at 260 nm''' | ||
[[Image:Screen_Shot_2013-10-05_at_5.21.54_AM.png]]<br> | |||
'''Adenosine Calibration Curve | '''Adenosine Calibration Curve at 260 nm | ||
''' | ''' | ||
== | [[Image:Screen_Shot_2013-10-05_at_4.31.20_AM.png]]<br> | ||
'''Class Calibration Curve for Adenosine at 259 nm''' | |||
[[Image:Screen_Shot_2013-10-05_at_4.43.15_AM.png]]<br> | |||
==Notes== | |||
Due to the time restrictions of the lab, the inosine spectra was taken the next lab (9/4/13). | Due to the time restrictions of the lab, the inosine spectra was taken the next lab (9/4/13). | ||
Grubb's test was conducted to determine the outliers in the pooled class data. | |||
<!-- ##### DO NOT edit below this line unless you know what you are doing. ##### --> | <!-- ##### DO NOT edit below this line unless you know what you are doing. ##### --> | ||
Revision as of 02:35, 5 October 2013
 Biomaterials Design Lab Biomaterials Design Lab
|
<html><img src="/images/9/94/Report.png" border="0" /></html> Main project page <html><img src="/images/c/c3/Resultset_previous.png" border="0" /></html>Previous entry<html> </html>Next entry<html><img src="/images/5/5c/Resultset_next.png" border="0" /></html> | ||||||||||||||||
|
The template for this lab can be seen from Dr. Hartings lab. Values are altered to accurately describe the lab that was conducted on this day. The template can be found here ObjectiveThe molar absorptivities of two different molecules, adenosine and inosine were determined in this lab using UV-Vis and Beer's law. The changes in UV-Vis spectra will be observed to determine changes in concentration of both adenosine and inosine. In order to do this, the molar absorptivity (ε) of both of these molecules will be known. A calibration curve from the class data will be created. From this data the standard deviation, Confidence Interval (90% and 95% confidence) will be calculated and Grubb's test will be performed to determine the outlier.
DilutionsStock Solutions Stock solutions were made to create these dilutions for each molecule. The calculations for the stock solution and dilutions were performed before lab.
The RANDOM concentration was 0.25*10^-5 for adenosine and 0.4*10^-5 for inosine. Preparation of Dilutions of Adenosine Preparation of Dilutions of Inosine
DataAdenosine Absorbance Spectrum at 260 nm Adenosine Calibration Curve at 260 nm Class Calibration Curve for Adenosine at 259 nm NotesDue to the time restrictions of the lab, the inosine spectra was taken the next lab (9/4/13). Grubb's test was conducted to determine the outliers in the pooled class data. | |||||||||||||||||