Molecular Recognition Laboratorium
Contact Information
Molecular Recognition Laboratorium,
Institute für medizinische Immunologie
CHARITÉ - UNIVERSITÄTSMEDIZIN BERLIN

Hessische Str. 3-4 D-10115 Berlin, Germany phone +49-30-450 524092 fax +49-30-450 524942 mail annette.hayungs@charite.de web charite.de
Group Leader
Group Members
- Bernhard Aÿ, Postdoc
- Prisca Boisguérin, Postdoc
- Zerrin Fidan, Doktorandin
- Annette Hayungs - Secretary
- Marc Hovestädt - Doktorand
- Ines Kretzschmar - CTA
- Christiane Landgraf - CTA
- Carsten Mahrenholz - Doktorand
- Judith Müller - Doktorandin
- Livia Otte - Postdoc
- Rolf-Dietrich Stigler - IT- u. Sicherheitsbeauftragter
- Víctor Tapia - Doktorand
- Julia Triebus - Diplomantin
- Lars Vouillème - Doktorand
- Eike Wolter - Diplomant
Research interest
Technological Development of the Peptide Array Technologies
by Victor Tapia
FIGURE
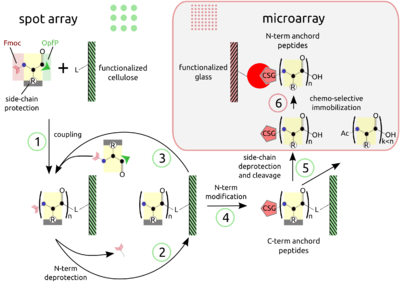
PEPTIDE ARRAYS The combination of SPOT peptide synthesis (steps 1 to 4) with appropriate immobilization techniques on glass supports (steps 5 and 6) is wide spread. The SPOT technology provides low-scale but high-throughput synthesis, while immobilization of pre-synthesized peptides offers the benefit of a "chemical" purification step and flexible array design. Additionally, the glass support is compatible with fluorescence detection and offers the possibility to miniaturize binding assays. Beyond economy, the later point is essential for quantitative measurements at the steady-state of binding activity, as has been described [Ekins 1998] and can be proven by the mass-action law.
The basic point of this technology is the simultaneous display of a systematic collection of peptides on a planar support, on which numerous bimolecular interaction assays can be carried out under homogeneous conditions.
Structural Modularity in Protein-Protein Recognition
by Victor Tapia
FIGURE
Protein Interaction Domains
The organisation of living systems is a complex network of molecular interactions. Proteins are a central component of the network as they may bind to other proteins as well as to phospholipids, nucleic acids and small molecules to interconnect the diverse physiological functions of the cell. In the background of these observations, the existence of a molecular recognition code for cellular organisation is very suggestive.
MORE TEXT
Structural analysis of functional protein complexes suggests at least two classes of protein-protein interaction that may be extendable to the other kinds of protein interactions. In the first class, the complementary surfaces of the interacting partners are both extensive. Under these circumstances, the residues involved in each interacting surface come together only upon protein folding. The second class consists on asymmetric interactions, where a protein domain (folding motive of moderate size like a pocket on the protein’s surface) may dock a short lineal peptide motive (a peptide ligand) on the partner protein. While interactions over extensive surfaces cannot be inferred, the binding determinants of a protein interaction domain (PID) may be mapped to short peptides matching the sequence of the ligand peptide. The importance of small recognition domains in the formation of protein complexes by binding to short lineal peptides was demonstrated in the late 1980s and early 1990s (Sadowski, I. et al., 1986; Ren, R. et al., 1993; Mayer, B.J. et al., 1993).
In this era of extensive genome sequencing, many PIDs have been discovered. The interaction partners and, therefore, the functions of such proteins may be determined by identifying the critical binding sites for one family member through evolutionary tracing (Lichtarge, O. et al., 1996) or through high-parallel screening of functional protein arrays (Phizicky, E. et al, 2003). Many of the PIDs in proteins can be grouped into families that show clear evidence of their evolution from a common ancestor, and genome sequences from Saccharomyces cerevisiae to Homo sapiens reveal large numbers of proteins that contain one or more common domains.
Src Homology Families - A PID Prototype
In a pioneering work on the kinase function and transforming activity of the Fujinami Sarcoma Virus, Sadowski et al. (1986) discovered “a unique domain… (which) is absent from kinases that span the plasma membrane” and concluded that “the presence of this noncatalytic domain in all known cytoplasmic tyrosine kinases of higher and lower eucaryotes argues for an important biological function... the noncatalytic domain may direct specific interactions of the enzymatic region with cellular components that regulate or mediate tyrosine kinase function”. These regions were called Src homology 2 (SH2) and 3 (SH3), the name SH1 being reserved to the catalytic region. Since then, the gained knowledge on SH domain function has been a paradigm in our understanding of PID biochemistry.
MORE TEXT
The structure of SH2 family members involves about 100 residues that, in the case of the kinase Src, are located N terminal to the catalytic region and resembles a pocket dominated by a β-sheet sandwiched between a pair of α-helices. SH2 domains bind the protein containing them to a second protein on a phosphorylated tyrosine residue (pY) in a specific amino acid sequence context (Ladbury, J.E. et al., 2000).
The SH3 domain structure, also found in cytoplasmic kinases like Src, consists largely of two sheets that form a partly open β-barrel. The ligand-binding site is a hydrophobic surface showing three shallow pockets or grooves defined by conserved aromatic residues. The ligand adopts an extended, left-handed helical conformation termed the polyproline-2 (or PPII) helix. Two of the binding pockets of the SH3 domain are occupied by two hydrophobic proline dipeptides on two adjacent turns of the helix, whereas the third ‘specificity’ pocket in most cases interacts with a basic residue in the ligand distal to the xPxxP core conserved motive of the PPII helix (Mayer, B.J. et al., 2001).
The amino acids located at the binding site for the phosphorylated polypeptide of SH2 and for the polyprolin core of SH3 have been the slowest to change during the long evolutionary process that produced the large SH2 and SH3 families of peptide recognition domains. Because mutation is a random process, this result is attributed to the preferential elimination during evolution of all organisms whose SH domains became altered in a way that inactivated the SH-binding site, thereby destroying the function of the SH domain. Are the PID/ligand interactions specific enough or must a certain interaction compete with the bulk of structurally similar structures in a struggle for dynamical complex formation?
From Promiscuous Recognition Events to Mutually Exclusive Cellular Responses
The elucidation of functional pathways of signal transduction, biochemical function or gene regulation, is firstly addressed in proteomics by deriving interaction networks depicting ideally all interactions in the cell. Several attempts have been done in this direction on different model organisms and with varied methods, including co-purification by affinity chromatography (Ho, Y. et al., 2002; Gavin, A.-C. et al., 2002; Bouwmeester, T. et al., 2004), yeast two-hybrid, phage display, spot synthesis, etc. A comparison of datasets derived by individual methods demonstrates that different methods have different potential. For example, affinity chromatographic approaches are biased to tight interactions such as those involving extensive complementary surfaces, while interactions in which one of the two partners contains at least one PID are more frequent in the two-hybrid database. The higher sensitivity of the so called synthetic approaches (yeast two-hybrid, phage display and spot synthesis) make them better suited for detecting PID-mediated interactions since their peptide affinity in terms of Kd falls in the 10 – 100 µM range (high Kd low affinity). However, this advantage is counterbalanced by a low specificity, especially of the yeast two-hybrid approach.
MORE TEXT
In order to correct this deficiency a double check-up of the information fed into the interaction databases is recommended. This can be achieved by deriving two interaction networks through orthogonal (fundamentally different) synthetic methods and then considering only the intersection between the two datasets (Tong, A.H.Y. et al.,2002; Castagnoli, L. et al., 2004; Landgraf, C. et al., 2004). False positive reports are thus reduced if the causes for measurement error are different in each method.
The strength of this combined approach to deliver physiologically relevant interactions has been proven for a phage display/yeast two-hybrid intersected dataset (Tong, A.H.Y. et al.,2002). A notable conclusion of this approach is that the intersected dataset of proteins that are able to interact with a given PID is larger than expected when cellular events are viewed as a precise wiring of the proteins in the cell. Although a set of these biochemically potential binders may have no physiological relevance due to expression at different times or tissues, in vitro disrupted structures, etc., the paradox of promiscuous recognition and mutually exclusive responses seems to be inherent to PID mediated interactions: the recent work of Landgraf et al. (2004) supports the observation that a large fraction of natural peptides with the biochemical potential to bind to any given SH3 domain is actually used in vivo to mediate the formation of a complex.
An additional difficulty to derive functional interaction pathways is that the difference in affinity between ‘specific’ and ‘non-specific’ interactions has been shown to be less than two orders of magnitude in the case of SH2 and its peptide ligands (Songyang, Z. et al., 2004). Even when granted that the recognition specificity of intact proteins by SH3 domains is greater than for SH3-peptide recognition, affinity is not raised above one order of magnitude (Arold, S. et al., 1998; Lee, C.H. et al., 1995). Moreover, the ability of a point-mutant Src SH2 domain to effectively substitute for the SH2 domain of the Sem-5 protein in activation of the Ras pathway in vivo emphasises that the specificity of Sh2-mediated interactions is not great (Marengere, L.E. et al., 1994). Consider the later statements under the light of the fact that the affinity of the protein OppA for its ligands is in the range of two orders of magnitude (Oppa is involved in the mopping of peptides in the bacterial periplasm exhibiting no sequence specificity). Tu put it all into a nutshell: the described facts lead to a view of large and promiscuous SH-mediated interaction networks.
Since it is possible to generate mutant SH3 domains that have up to 40-fold higher affinity than their wild-types (Hiipakka et al, 1999) , the potential of these domains as research tools and as source of lead compounds for pharmaceutical development can not be overseen. Furthermore, a question cannot be overheard in our minds: which is the functional advantage of maintaining relative low affinity and selectivity for PID-mediated interactions, instead of optimizing the potential affinity of PIDs? And further: how can PID-dependent interaction pathways achieve precise cellular responses?
A comfortable view is that sufficient effective selectivity can be brought by compartmentalization, additive effects of multiple separate interactions, cooperative assembly of multiprotein complexes, etc. and that this effects can sustain linear functional pathways. An interesting insight into this mater has been advanced by Zarrinpar, A. et al. In their work (2003) the authors find out that while metazoan SH3 domains may rescue the functionality of mutated Sho1-SH3 of the yeast, in the set of yeast-own SH3 domains this promiscuity is forbidden. They thus conclude and confirm that, on the background of diverging SH3 domains, negative selection has drifted the ligand sequences to non-overlapping areas of the particular SH3 binding regions on the sequence space (please see Figure 1 and section 3.2 for further explanation of the concept). Alternatively, a divergence process of the shape of the binding region may be ‘guided’ by positive selection to avoid overlapping and, thus, promiscuitive interactions. Nevertheless, the picture of linear functional pathways is being revolutionized by a more probabilistic view of a dynamical equilibrium between multiple interactions, in which “the central organizing principle is a vast and ever-shifting web of interactions, from which output is gauged by global changes in complex binding equilibria” (Mayer, B. J., 2001).
Founding
Who's visiting
start on 16.03.2009
<html> <a href="http://clustrmaps.com/counter/maps.php?url=http://openwetware.org/wiki/Molecular_Recognition_Laboratorium" id="clustrMapsLink"><img src="http://clustrmaps.com/counter/index2.php?url=http://openwetware.org/wiki/Molecular_Recognition_Laboratorium" border=1 alt="Locations of visitors to this page"onError="this.onError=null; this.src='http://www.meetomatic.com/images/clustrmaps-back-soon.jpg'; document.getElementById('clustrMapsLink').href='http://clustrmaps.com/'"> </a> </html>
Science in the city: A small series about the everyday life of scientists in berlin
FOLLOW ME >>>>