Maloof Lab:Jose M. Jimenez-Gomez: Difference between revisions
mNo edit summary |
mNo edit summary |
||
| Line 22: | Line 22: | ||
---- | ---- | ||
<br> | <br> | ||
It is well known that plants from different light environments exhibit different degrees of responsiveness to similar light stimulus. | It is well known that plants from different light environments exhibit different degrees of responsiveness to similar light stimulus. For example, plants accommodated to sunny environments detect foliar shade from neighboring vegetation and respond increasing their petioles/stems and reducing the time to reproduction, a phenomenon called the "shade avoidance response". On the other hand, plants adapted to live under dense canopies are less sensitive to the shade and present a reduced shade avoidance response. | ||
To identify the molecular mechanisms underlying this differences we are performing QTL analysis using a previously developed, well characterized Recombinant Inbred Line set descent from two different natural populations of <i>Arabidopsis thaliana</i>: Bayreuth, originary from the German low altitude fallow lands, and Shahdara, from the high mountains of Tadjikistan <cite>Loudet02</cite>.<br> | To identify the molecular mechanisms underlying this differences we are performing QTL analysis using a previously developed, well characterized Recombinant Inbred Line set descent from two different natural populations of <i>Arabidopsis thaliana</i>: Bayreuth, originary from the German low altitude fallow lands, and Shahdara, from the high mountains of Tadjikistan <cite>Loudet02</cite>.<br> | ||
We grew replicated individual RILs in environments simulating shade and sun conditions and | We grew replicated individual RILs in environments simulating shade and sun conditions and characterized them on a number of traits associated with the shade avoidance response syndrome. For the QTL analysis we calculated a shade avoidance response index fitting fixed effect models to the phenotipic data, and used an available genetic map for the population that includes more than 500 Single Feature Polymorphism (SFP) markers <cite>West06</cite>.<br> | ||
<br> | <br> | ||
<br> | <br> | ||
| Line 35: | Line 35: | ||
<br> | <br> | ||
<br> | <br> | ||
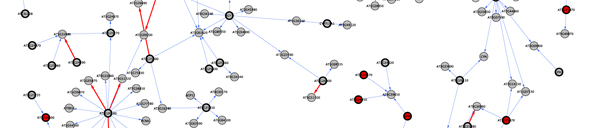
We are focusing | We are now focusing in a chromosomal region containing about 200 genes to fine map and identify the gene responsible for the differential response to shade between the two natural populations. To do this we employ traditional genetic approaches as well as genomic and network analysis. We are developing a protocol to construct gene networks that will help us consider candidate genes based on coexpression with other genes across microarray experiments <cite>Riken</cite>, colocalization with expression QTLs <cite>West07</cite>, functional categorization <cite>GO_Classification</cite> and presence of polymorphisms between the parental lines <cite>Clark07</cite>. | ||
<br> | <br> | ||
<br> | <br> | ||
| Line 48: | Line 48: | ||
---- | ---- | ||
<br> | <br> | ||
We use bioinformatics to detect Single Nucleotide Polymorphisms among the numerous tomato EST sequences available in the databases. | We use bioinformatics to detect Single Nucleotide Polymorphisms among the numerous tomato EST sequences available in the databases. We aim to estimate divergence rates over large regions of the genome of selected species, and obtain a new set of molecular markers useful in natural variation studies. We use this data to perform functional and evolutionary pre-genomic analyses on this dataset, which should give us an idea of which gene families evolve more rapidly/slowly and have been important during tomato domestication. | ||
<br> | <br> | ||
<br> | <br> | ||
| Line 60: | Line 60: | ||
|- | |- | ||
|align="center"|[[Image:PHYB_alignment.jpg|center]] | |align="center"|[[Image:PHYB_alignment.jpg|center]] | ||
<small>amino-acid changes in a fragment of the PHYB gene in 8 | <small>amino-acid changes in a fragment of the PHYB gene in 8 species, red and black bars indicate non-conserved/conserved amino-acid changes respectively</small> | ||
|- | |- | ||
|} | |} | ||
Revision as of 20:09, 11 November 2008
|
Room 2115 |
QTL analysis of the shade avoidance response in Arabidopsis
Single Nuncleotide Polymorphism discovery between wild Tomato species
Molecular evolution of PHYTOCHROME B
Proteomics of light perception
References
|