IGEM:Harvard/2006/Adaptamers/Notebook/2006-8-28: Difference between revisions
No edit summary |
|||
| Line 48: | Line 48: | ||
[[Image:cst828-displace5washesgraph.jpg]] | [[Image:cst828-displace5washesgraph.jpg]] | ||
This data confirmed much of what was seen for the same experiment on 8-26. while strand displacement successfully quenches adaptamer function, increasing amounts of displacement strand led to increased fluorescence. Most interestingly, samples containing displacement strands retained fluorescence following numerous washes in comparison to samples of only adaptamers. I have proposed a mechanism explaining this observation as follows: | |||
===Lever mechanism=== | |||
Thrombin is represented by a rectangle, streptavidin by an oval. The double-helix regions of our adaptamers in effect create long, rigid rods (A). The longer these rods are, the more susceptible they are to washes since applying pressure toward the thrombin side of the adaptamer (see arrow in A) results in a large amount of mechanical advantage that enables the dissociation of the adaptamer from the surface of the streptavidin bead. This explains the lack of fluorescence observed using the adaptamer with the longest linking region, A50, despite Western Blot results. In contrast, addition of displacement strand (red) reduces the length of any rigid regions, bringing about flexibility that makes it less susceptible to washes (B). Moreover, displacement strands provide multiple attachment sites for T20, increasing the possibility of attaching multiple thrombins to a single streptavidin molecule. | |||
[[Image:LeverMechanism.jpg]] | |||
Latest revision as of 18:23, 29 October 2006
8/28
Strand Displacement round 2
General setup = 40 pmol A20 + 100 uL Streptavidin magnetic beads + 160 pmol thrombin
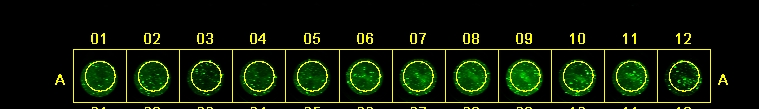
| Well | Condition |
| 1 | general setup |
| 2 | general setup |
| 3 | adaptamer = A50 |
| 4 | 10 pmol R20.20 added |
| 5 | 20 pmol R20.20 added |
| 6 | 40 pmol R20.20 added |
| 7 | 10 pmol R20.35 added |
| 8 | 20 pmol R20.35 added |
| 9 | 40 pmol R20.35 added |
| 10 | 40 pmol T20 instead of A20 |
| 11 | 40 pmol S20 instead of A20 |
| 12 | no thrombin added |
- After 3 washes
Image taken at intensity 3
- After 5 washes
This data confirmed much of what was seen for the same experiment on 8-26. while strand displacement successfully quenches adaptamer function, increasing amounts of displacement strand led to increased fluorescence. Most interestingly, samples containing displacement strands retained fluorescence following numerous washes in comparison to samples of only adaptamers. I have proposed a mechanism explaining this observation as follows:
Lever mechanism
Thrombin is represented by a rectangle, streptavidin by an oval. The double-helix regions of our adaptamers in effect create long, rigid rods (A). The longer these rods are, the more susceptible they are to washes since applying pressure toward the thrombin side of the adaptamer (see arrow in A) results in a large amount of mechanical advantage that enables the dissociation of the adaptamer from the surface of the streptavidin bead. This explains the lack of fluorescence observed using the adaptamer with the longest linking region, A50, despite Western Blot results. In contrast, addition of displacement strand (red) reduces the length of any rigid regions, bringing about flexibility that makes it less susceptible to washes (B). Moreover, displacement strands provide multiple attachment sites for T20, increasing the possibility of attaching multiple thrombins to a single streptavidin molecule.