Frankel:Force Spectroscopy: Difference between revisions
No edit summary |
No edit summary |
||
| (24 intermediate revisions by the same user not shown) | |||
| Line 1: | Line 1: | ||
{{Frankel}} | {{Frankel}} | ||
<br> | |||
'''<font color=#000000 font size=8>Force Spectroscopy</font>''' | '''<font align="center" font color=#ffffff font size=8>______ </font>''''''<font align="center" font color=#000000 font size=8>Force Spectroscopy</font>''' | ||
{| cellspacing=" | {| cellspacing="3px" | ||
<div> | |||
<div | |||
| align="center" width=400px style="border: 2px solid #000033; background-color:#FFFFFF; padding:1em;" class="plainlinks" valign="top"| | |||
[[Image:Spectroscopy_complete.jpg| | |||
'''<font color=#000000 font size=3>Picking up and unfolding a single protein</font>''' | |||
[[Image:Spectroscopy_complete.jpg|350px]] | |||
</div> | </div> | ||
| align=" | | align="center" rowspan="2" width=21px style="border: 2px solid #000033; background-color:#ffffff; padding:1em;" valign="top"| | ||
'''<font color=#FFFFFF font size=3>Assembly</font>''' | '''<font color=#FFFFFF font size=3>Assembly</font>''' | ||
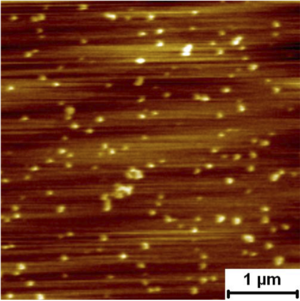
'''<font color=#000000 font size=3> | '''<font color=#000000 font size=3>Nano-mechanics of HIV GP160</font>''' | ||
[[Image:GP160mica.png|300px]] | [[Image:GP160mica.png|300px]] | ||
[[Image: | [[Image:GP_160.png|300px]] | ||
[[Image: | |||
[[Image: | '''<font color=#FFFFFF font size=6>GP160mica</font>''' | ||
[[Image:Stlow.png|330px]] | |||
[[Image:Hislow.png|330px]] | |||
---- | ---- | ||
'''<font color=#000000 font size=3> | '''<font color=#000000 font size=3 font align="justify"> | ||
Typical sawtooth unfolding pattern of a single molecule of the HIV virion protein GP160. Unfolding forces can be extracted from the sawtooth pattern by applying the worm like chain model.</font>''' | |||
{|width="*" | {|width="*" | ||
| | | | ||
{| style="width: 728px; text-align:center;font-size:12px;font-variant: small-caps;width: 18px; " align=" | {| style="width: 728px; text-align:center;font-size:12px;font-variant: small-caps;width: 18px; " align="justify" | ||
|- | |- | ||
| Line 53: | Line 58: | ||
|} | |} | ||
|- | |- | ||
| align=" | | align="center" width=21px style="border: 2px solid #000000; background-color:#FFFFFF; padding:1em;" valign="top"| | ||
'''<font color=# | '''<font color=#000000 font size=3> Unfolding the ECM protein fibronectin </font>''' | ||
[[Image:FNsawt.png|300px]] | [[Image:FNsawt.png|300px]] | ||
[[Image:HisFN.png|300px]] | [[Image:HisFN.png|300px]] | ||
| Line 65: | Line 69: | ||
---- | ---- | ||
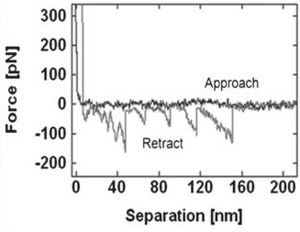
'''<font | '''<font align="justify" font size=3> | ||
Sawtooth pattern on the retraction force curve indicating the unfolding of fibronectin. The average rupture force distribution of the protein on mica surface was 85.1 ± 2.7 pN. | |||
Latest revision as of 18:22, 9 January 2013
<owwmenu align="center" font="helvetica" bold="1" color="white" bgcolor="black" hovercolor="black" bghovercolor="orange" topfontsize="10" fontSize="10" image="Danbanner-bio-machines.jpg" >
Home=Frankel
Members=#,Principal Investigator=Frankel:Lab_Members, PhD students=Frankel:Lab_Members, Alumni=Frankel:Lab_Members
Contact=Frankel:Contact
Collaborators=Frankel:Collaborators
Publications=Frankel:Publications
Lab=Frankel:Research
Research=#,Force Spectroscopy=Frankel:Force Spectroscopy,HIV/Virus=Frankel:HIV/Virus,ECM Proteins=Frankel:ECM Proteins,Cyberplasm=Frankel:Cyberplasm,Cancer=Frankel:Cancer
'______ 'Force Spectroscopy
|
Picking up and unfolding a single protein |
Assembly Nano-mechanics of HIV GP160 GP160mica
Typical sawtooth unfolding pattern of a single molecule of the HIV virion protein GP160. Unfolding forces can be extracted from the sawtooth pattern by applying the worm like chain model.
| |
|
Unfolding the ECM protein fibronectin
Sawtooth pattern on the retraction force curve indicating the unfolding of fibronectin. The average rupture force distribution of the protein on mica surface was 85.1 ± 2.7 pN. |