Biomolecular Breadboards:Preliminary Data
| Home | Protocols | DNA parts | Preliminary Data | Models | More Info |
This page contains some data that we have taken with the TX-TL breadboard.
Plasmid Expression of GFP
Using pBEST-OR2-OR1-Pr-UTR1-deGFP-T500, a plasmid enhanced for GFP expression, the biomolecular breadboard is able to express mass at equal concentrations to comparable bacteriophage in-vitro systems (J. Shin and V. Noireaux, 2010).
Expression of plasmids can be optimized by concentration.
|
Figure 1. eGFP expression as a function of plasmid DNA template. Plasmid DNA pBEST-OR2-OR1-Pr-UTR1-eGFP-T500 is varied by concentration.
Protecting Linear DNA from Exonuclease-Mediated Degradation
Current standards for circuit design utilize plasmids for DNA template, which require time-consuming subcloning steps. However, circuits based on linear DNA require only PCR assembly or gene synthesis, which drastically decreases preparation time. As a purely extract-derived system, our biomolecular breadboard exhibits exonuclease activity which degrades linear DNA. We are developing multiple technologies to protect linear DNA from exonuclease degradation. These include:
- Protecting linear DNA using noncoding segments
- Inhibiting RecBCD exonuclease with gamS
- Adding thiosulfate bonds to 5' ends
Protection Sequences
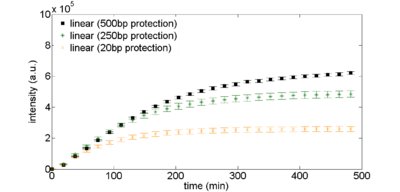
Figure 2: Exonuclease protection using non-coding DNA. Linear DNA templates, 2nM, are derived from plasmid DNA pBEST-OR2-OR1-Pr-UTR1-eGFP-Del6-229-T500 with varying amounts of noncoding DNA surrounding the coding sequence.
GamS


Figure 3. (left) GamS from lambda phage, an inhibitor of RecBCD complex, inhibits template DNA degradation. Linear DNA templates, 2nM, are derived from plasmid DNA pBEST-OR2-OR1-Pr-UTR1-eGFP-Del6-229-T500. GamS supplied at 3uM concentration. (right) Simulation results based on a simple ODE model.

Figure 4. eGFP expression as a function of linear DNA template, with gamS. Plasmid DNA pBEST-OR2-OR1-Pr-UTR1-eGFP-Del6-229-T500 is varied by concentration. GamS supplied at 3uM concentration.
Thiosulfate Bonds

Figure 5. Thiosulfate bonds protect against 5' exonuclease degradation. Linear DNA templates, 2nM, are derived from plasmid DNA pBEST-OR2-OR1-Pr-UTR1-eGFP-Del6-229-T500. 5 thiosulfate bonds are present at each 5' end. GamS supplied at 3uM concentration.
Protein Degradation
Implementing protein degradation is integral to enabling the function of many dynamical circuits, such as oscillators or feed-forward loops. We have been exploring the overexpression of AAA ATPases ClpXP to selectively target proteins with degradation tags. Initial experiments indicate that ClpXP, when overexpressed, can accelerate degradation of purified eGFP-ssrA (Fig. 1). We will further characterize this AAA ATPase family, as well as try to demonstrate its use through an incoherent feed-forward loop. We have not been able to replicate the results by adding purified ClpXP protein. However, we intend to express ClpXP off of plasmids or to create custom extract with ClpXP already overexpressed to enable rapid protein degradation.
