Biomolecular Breadboards:Preliminary Data: Difference between revisions
(→GamS) |
|||
| Line 31: | Line 31: | ||
[[Image:ts-1.png|400px]] | [[Image:ts-1.png|400px]] | ||
'''Figure X. Thiosulfate bonds protect against 5' exonuclease degradation.''' Linear DNA templates, 2nM, are derived from plasmid DNA pBEST-OR2-OR1-Pr-UTR1-eGFP-Del6-229-T500. 5 thiosulfate bonds are present at each 5' end. GamS supplied at 3uM concentration. | |||
Revision as of 23:55, 11 July 2012
| Home | Protocols | DNA parts | Preliminary Data | Models | More Info |
Preliminary Data
Plasmid Expression of GFP
Using pBEST-OR2-OR1-Pr-UTR1-eGFP-Del6-229-T500, a plasmid enhanced for GFP expression, the biomolecular breadboard is able to express mass at equal concentration to comparable bacteriophage in-vitro systems (J. Shin and V. Noireaux, 2010).
Expression of plasmids can be optimized by concentration.
Figure X. eGFP expression as a function of plasmid DNA template. Plasmid DNA pBEST-OR2-OR1-Pr-UTR1-eGFP-Del6-229-T500 is varied by concentration.
Protecting Linear DNA from Exonuclease-Mediated Degradation
Current standards for circuit design utilize plasmids for DNA template, which require time-consuming subcloning steps. However, circuits based on linear DNA require only PCR assembly or gene synthesis, which drastically decreases preparation time. As a purely extract-derived system, our biomolecular breadboard exhibits exonuclease activity which degrades linear DNA. We are developing multiple technologies to protect linear DNA from exonuclease degradation.
Protection Sequences
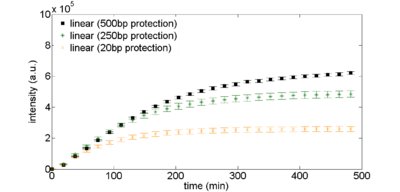
Figure 1: Exonuclease protection using non-coding DNA. Linear DNA templates, 2nM, are derived from plasmid DNA pBEST-OR2-OR1-Pr-UTR1-eGFP-Del6-229-T500 with varying amounts of noncoding DNA surrounding the coding sequence.
GamS

Figure X. GamS from lambda phage, an inhibitor of RecBCD complex, inhibits template DNA degradation. Linear DNA templates, 2nM, are derived from plasmid DNA pBEST-OR2-OR1-Pr-UTR1-eGFP-Del6-229-T500. GamS supplied at 3uM concentration.

Figure X. eGFP expression as a function of linear DNA template, with gamS. Plasmid DNA pBEST-OR2-OR1-Pr-UTR1-eGFP-Del6-229-T500 is varied by concentration. GamS supplied at 3uM concentration.
Thiosulfate Bonds
 Figure X. Thiosulfate bonds protect against 5' exonuclease degradation. Linear DNA templates, 2nM, are derived from plasmid DNA pBEST-OR2-OR1-Pr-UTR1-eGFP-Del6-229-T500. 5 thiosulfate bonds are present at each 5' end. GamS supplied at 3uM concentration.
Figure X. Thiosulfate bonds protect against 5' exonuclease degradation. Linear DNA templates, 2nM, are derived from plasmid DNA pBEST-OR2-OR1-Pr-UTR1-eGFP-Del6-229-T500. 5 thiosulfate bonds are present at each 5' end. GamS supplied at 3uM concentration.
