BME103:T930 Group 6 l2: Difference between revisions
No edit summary |
|||
| Line 199: | Line 199: | ||
<!--- A description of the diseases and their associated SNP's (include the database reference number and web link) ---> | <!--- A description of the diseases and their associated SNP's (include the database reference number and web link) ---> | ||
The SNP chosen for the lab is linked with Alzheimer's disease. Alzheimer's disease is the most common form of dementia. It has no cure and worsens as it progresses. Dementia is a loss of brain function and | The SNP chosen for the lab is linked with Alzheimer's disease. Alzheimer's disease is the most common form of dementia. It has no cure and worsens as it progresses. Dementia is a loss of brain function and Alzheimer's disease affects memory, thinking, and behavior. The associated SNP's for an increase in risk of Alzheimer's disease is rs1050283, which is located on the 12 chromosome. Information available from the NCBI website: | ||
http://www.ncbi.nlm.nih.gov/snp?term=rs1050283 | http://www.ncbi.nlm.nih.gov/snp?term=rs1050283 | ||
| Line 207: | Line 207: | ||
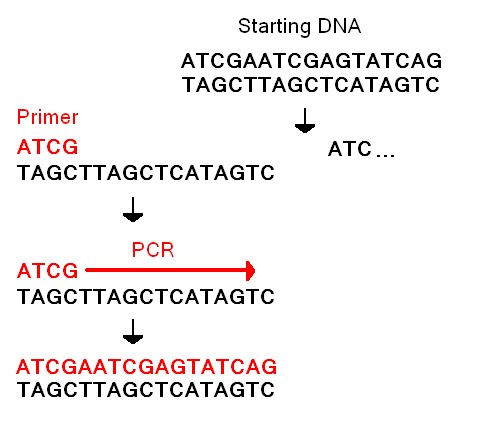
<!--- Include the sequences of your forward and reverse primers. Explain why a disease allele will give a PCR product and the non-disease allele will not. ---> | <!--- Include the sequences of your forward and reverse primers. Explain why a disease allele will give a PCR product and the non-disease allele will not. ---> | ||
The sequence that is connected with an increase risk of Alzheimer's disease is GGCTGGGCOCGGACATGGAGGACGTG[C/T]GCGGCCGCCTGGTGCAGTACCGCGG. | The sequence that is connected with an increase risk of Alzheimer's disease is GGCTGGGCOCGGACATGGAGGACGTG[C/T]GCGGCCGCCTGGTGCAGTACCGCGG. An increase in the risk for Alzheimer's disease comes from the base change C to a T. | ||
The reverse primer would be GGAGGACGTG[T]GCGGCCGCCT and the forward primer would be CCTCCTGCAG[A]CGCCGGCGGA. | The reverse primer would be GGAGGACGTG[T]GCGGCCGCCT and the forward primer would be CCTCCTGCAG[A]CGCCGGCGGA. | ||
A diseased allele will produce a PCR product because the gene associated with an increase risk that is trying to be tested for will have a T base in place of the normal C base. If the gene has this change in bases then the primers will be able to attach to the strands of DNA because the primers are designed to only attach to that specific sequence. | A diseased allele will produce a PCR product because the gene associated with an increase risk that is trying to be tested for will have a T base in place of the normal C base. If the gene has this change in bases then the primers will be able to attach to the strands of DNA because the primers are designed to only attach to that specific sequence. | ||
Revision as of 16:37, 28 November 2012
| Home People Lab Write-Up 1 Lab Write-Up 2 Lab Write-Up 3 Course Logistics For Instructors Photos Wiki Editing Help | |||||||||||||||||||||||||||||||||||
OUR TEAMLAB 2 WRITE-UPThermal Cycler Engineering System Design  Key Features To begin with we will change the frame from a wooden construction to recycled aluminum. This will maintain the light-weight portability of the machine, but will greatly reduce the fire hazard of heating a wooden box to the required temperatures. We will also add an additional row and column to the heating block to increase the product output without too much change to the overall size or cost of the machine. We will also add two carrying handles to either side of the machine. Because the machine is somewhat awkward to carry and hold the handles will greatly reduce the chances of dropping if they are properly used.  Instructions
ProtocolsMaterials
1. Using a micro-pipette, transfer 1.0-1.5 micro liters of desired DNA sample into at least three test tubes. 2. Micro-pipette approximately 3.0 micro liters of reagent solution, including forward primer, reverse primer, GoTaq polymerase and buffer solution, into each of the test tubes with the DNA. 3. Invert tubes, then turn upright to mix solution. 4. Plug in OpenPCR Machine and turn it on. 3. Open lid and place tubes into holder in PCR machine (can fit up to 25 tubes). Close the lid. 4. Program the folowing cycles on the OpenPCR program: - Stage 1: 1 cycle, 95 degrees Celsius for 3 minutes - Stage 2: 30 cycles, 95 degrees for 30 seconds, 57 degrees for 30 seconds, 72 degrees for 30 second - Stage 3: 72 degrees for 3 minutes - Final Hold: 4 degrees 5. Run reaction. 6. After cycles are complete, open the lid and remove tubes for signal reading (see DNA Measurement Protocol below). DNA Measurement Protocol 1. Remove lid from black box and invert the box, open the front flap. 2. Place one glass slide into the holder. 3. Depending on the amount of samples being used, take a clean pipette for each sample and label it so that it is only used for that sample of DNA. For example, label a pipette for SYBR green with a horizontal line on the pipette bulb, and label a pipette for diluted water with a vertical line on the bulb. Do NOT use pipettes for any other solution than what they are labelled for. 4. Using the designated pipette, squeeze two drops of SYBR green dye inside the round glass windows on slide, preferably the second dot of the second row. 5. Begin by adding a sample of diluted water to the SYBR green dye, and using the appropriate labelled pipette, add two drops in the same spot as the SYBR green dye. 6. Align the sample so the blue LED shines directly though it focusing the light on the other side. 7. Move holder apparatus inside the box so that no light reaches it. 8. Place smartphone with the flash off onto holder and direct camera lens at the slide apparatus. 9. Take photo with smartphone. 10. E-mail the taken image to computer. 11. On the computer open the image file using ImageJ (program can be downloaded from the Internet.) 12. In ImageJ, under the "analyze" tool bar select "Set Measurements." 13. Make sure the following boxes are checked, and no other boxes other than the ones listed: -Area -Mean Grey Value -Integrated Density 14. From the download folder drag the image file into ImageJ. 15. When the image is opened in ImageJ from the top drop down menu select "Image" and then "Color" and then "Split Channels." 16. Close out the red and blue channel, only use the green channel 17. With the Oval tool select only the entire drop of liquid, avoiding the background but encompassing every bit of the droplet. 18. After the droplet part of the image is selected, go to "Analyze" then "Measure" or press Ctrl + M. 19. Now move the circle to the background of the image by clicking and dragging on the circle, be careful not to change its size. 20. Save the results as an excel spreadsheet (set by default). 21. Once the Image has been processed, using a clean pipette, remove the sample of diluted water and SYBR green dye from the slide. 22. Repeat steps 4-21 using included calf thymus DNA as a control, and move back on the glass slide two rows to eliminate contamination. 23. Once the control has been analyzed, repeat step 4-21 using the sample DNA, ensuring to move back two rows on the glass slide each time a new sample is tested. 24. Once there are no more rows on the glass slide, carefully discard the used slide and begin using a new one. Research and DevelopmentBackground on Disease Markers
| |||||||||||||||||||||||||||||||||||