Arking:JCAOligoTutorial3
Basic cloning: How to implement the Construction File
This tutorial describes what we'll be doing in the lab as a demonstration of basic cloning. This procedure describes what you'll be doing and seeing during cloning. There's no quiz at the end, but you'll have plenty of real-life experience with this procedure. First of all, let's examine the construction file:
Construction of GFP Biobrick 2.0 basic part PCR ca1123F/ca1123R on pSB1A2-I13522 (748 bp, BglII/XhoI/DpnI) Sub into pBca1102 (BglII/XhoI, 2159+697, L) Product is pBca1102-Bca1123 ---------------------------------------- ca1123F Forward BglII for eXtreme GFP basic part ctctgAGATCTatgcgtaaaggagaagaac ca1123R Reverse XhoI for eXtreme GFP basic part gcaaaCTCGAGttaGGATCCttatttgtatagttcatccatgc
Go ahead and download all the files and simulate it in your editor to confirm that it "works":
Media:JCAseq_pSB1A2-I13522.str Media:JCAseq_pBca1102.str Media:JCAseq_pBca1102-Bca1123.str
In this construction, we are using a plasmid pBca1102 I call "ORF expressor." This plasmid holds open reading frame basic parts, but it places a constitutive (meaning always on) promoter and ribosome binding site upstream of the Biobrick BglII site. The result of this is that the open reading frame will make the encoded protein when the plasmid is inserted into E. coli cells. However, it is still a "basic part" in the sense that the ORF is still flanked by BamHI and BglII restriction sites. In this construction file, we start with the parent plasmid pBca1102 which has an RFP gene inside these restriction sites. Cells with pBca1102 are therefore red. When we digest with BglII/XhoI, we remove the RFP gene. We'll paste in the PCR-amplified GFP gene, and upon insertion of the new plasmid into E. coli, the cells should be green. So, that's the game here, now let's see how specifically to do it.
Step 1: The PCR
When you receive oligos from IDT, it comes as a liophylized solid (ie, a dry powder). You must resuspend it in water to a final concentration of 100uM (uM is 10^-6 molar). Fortunately, IDT measures how many moles of DNA are present in the tube and writes it on the tube. Let's say the tube says it's 23.4nmoles of material. You would therefore add 234uL of water to the tube. I leave it to you to do the math and confirm that. To set up the PCR, you'll want to use 10uM stocks, so do a 10x dilution of the oligos. To achieve this, add 1uL of 100uM oligo to 9uL of water, then mix it by tapping it on the bench. Now you can set up the PCR.
A general procedure for setting up a PCR reaction is to make the following cocktail:
39 uL water 5 uL 2 mM in each dNTPs (meaning a mixture of 2mM dATP, 2mM dCTP, 2mM dTTP, and 2mM dGTP) 5 uL 10x Expand Buffer "2" 0.75 uL of 0.75 uL Expand Polymerase "1" 1 uL of 10uM Oligo 1 1 uL of 10uM Oligo 2 0.5 uL of template DNA
Add the ingredients in order. In general, never add an enzyme to a reaction until after you've added the buffer. The reason for this is that proteins can denature if they are diluted into a solution that is not buffered. It doesn't matter when you add the oligos and template, though.
Now mix up the sample by tapping and place it in the thermocycler. Choose the temperature program based on the size of the fragment:
PCR product Size Program Under 1kb 55 1kb to 2kb 2K55 2kb to 4kb 4K55 over 4kb 8K55
These programs principly differ in the length of time for each extension step during the cycles. This is necessary because longer PCR products take more time to polymerize. In general, it is OK to run a longer program than is necessary for the PCR product, but going shorter will cause failure. Sometimes doing longer extension times than are necessary will lead to aberrant products, so you should make an effort to use the right program. Alright, run the program.
Now you want to run an analytical gel to make sure you got the right product. There is another kind of gel you'll run later called a "preparative" gel. The distinction is that the analytical gel is run simply to examine the products. The prepartive gel is done to separate DNA fragments, and bands are individually excised from the gel and purified. You will add 5uL of 10x loading buffer to the PCR reaction, and then load 5 uL of material to a well of the gel. In another lane, add 5uL of molecular weight marker. You'll use the marker to estimate the size of your PCR band. Below is an example of a PCR product.
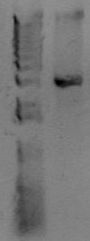

At the right of the PCR product is the known sizes present in the Hi-Lo molecular weight markers. The size of this PCR product is therefore somewhere around 3kb. After running this gel, estimate the size of your PCR product. Does it match the size you predicted for the construction file? Also, look at the quality of the band. Is it a single band or multiple bands? Is it tight or kindov fuzzy? If any of that fails, you'll need to troubleshoot