Abhishek Tiwari:SOFTWARE NOTES
SOFTWARE NOTES
<html> <a href="http://itol.embl.de/"><img src="http://itol.embl.de/img/shot.jpg" alt="iTOL" border="0"></a>
</html>
Oxford Bioinformatics Volume 23 | Number 1 | 1 January 2007
- Interactive Tree Of Life (iTOL): an online tool for phylogenetic tree display and annotation
Synopsis
iTOL or Interactive Tree Of Life is an online tool for the display and manipulation of phylogenetic trees. iTOL uses Shockwave Flash and Javascript to display trees. Trees can be interactively pruned and re-rooted. Various types of data such as genome sizes or protein domain repertoires can be mapped onto the tree. iTOL can display phylogenetic trees in either normal or circular mode. Branch lenghts and bootstrap values can be provided. Trees can have up to 5 datasets associated and displayed. iTOL's branch pruning, rerooting and collapsing functions allow you to customize the tree however you like. It also support Swapping of sub-branches along with Branch and leaf popups. It is a really handy tool with a lot features. ()
BMC Bioinformatics Volume 7 | MARCH 2006
- The Gaggle: An open-source software system for integrating bioinformatics software and data sources
Synopsis
Gaggle[1]-a simple, open-source Java software environment that helps to solve the problem of software and database integration. Guided by the classic software engineering strategy of separation of concerns and a policy of semantic flexibility, it integrates existing popular programs and web resources into a user-friendly, easily-extended environment.The Gaggle uses Java RMI and Java Web Start technologies.In the Gaggle, semantic flexibility – the notion that "word meanings are not ... fixed and unchanging, but tend to vary according to the context of their use" – is seen as a solution to the complications of data integration, rather than as a problem that must be solved before integration can begin. Four data types (names, matrices, networks, and associative arrays), distilled into a semantically simple form, are passed between the geese, whereupon they take on richer meaning in the context of each goose.
Oxford Bioinformatics Volume 22 | Number 22 | 15 November 2006
- Helix Interaction Tool (HIT): a web-based tool for analysis of helix-helix interactions in proteins
Synopsis
In many proteins, helix–helix interactions can be critical to establishing protein conformation (folding) and dynamics, as well as determining associations between protein units. However, the determination of a set of rules that guide helix–helix interaction has been elusive. In order to gain further insight into the helix–helix interface, we have developed a comprehensive package of tools for analyzing helix–helix packing in proteins. These tools are available at http://helix.gersteinlab.org. They include quantitative measures of the helix interaction surface area and helix crossing angle, as well as several methods for visualizing the helical interaction. These methods can be used for analysis of individual protein conformations or to gain insight into dynamic changes in helix interactions. For the latter purpose, a direct interface from entries in the Molecular Motions Database to the HIT site has been provided.
TreeDyn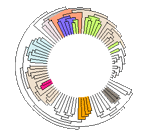 http://www.treedyn.org/
http://www.treedyn.org/
BMC Bioinformatics Volume 7 | OCTOBER 2006
- TreeDyn: towards dynamic graphics and annotations for analyses of trees
Synopsis TreeDyn is a tree visualization and annotation tool which includes tools for tree manipulation and annotation and uses meta-information through dynamic graphical operators or scripting to help analyses and annotations of single trees or tree collections.TreeDyn is using annotations and dynamic graphical methods for editing and analyzing multiple trees. The main features of TreeDyn are 1) the management of multiple windows and multiple trees per window, 2) the export of graphics to several standard file formats with or without HTML encapsulation and a new format called TGF, which enables saving and restoring graphical analysis, 3) the projection of texts or symbols facing leaf labels or linked to nodes, through manual pasting or by using annotation files, 4) the highlight of graphical elements after querying leaf labels (or annotations) or by selection of graphical elements and information extraction, 5) the highlight of targeted trees according to a source tree browsed by the user, 6) powerful scripts for automating repetitive graphical tasks, 7) a command line interpreter enabling the use of TreeDyn through CGI scripts for online building of trees, 8) the inclusion of a library of packages dedicated to specific research fields involving trees.
