20.109(S13):PCR and paper discussion (Day3): Difference between revisions
| Line 28: | Line 28: | ||
===Part 1: Prepare PCR to detect bacterial 16S=== | ===Part 1: Prepare PCR to detect bacterial 16S=== | ||
#Begin by carefully labeling each PCR tube that you will use with the date, sample name, and a unique symbol and/or color for quick identification. Filling in the cap tab with your team color will usually suffice. | #Begin by carefully labeling each PCR tube that you will use with the date, sample name, and a unique symbol and/or color for quick identification. Filling in the cap tab with your team color will usually suffice. | ||
| Line 56: | Line 55: | ||
| | | | ||
|- | |- | ||
| | | 5% BSA (100X stock) | ||
| 0.5 | | 0.5 | ||
| | | | ||
Revision as of 19:34, 8 February 2013
Introduction
Last time you prepared a DNA pool from a bird stool sample, and today you will amplify 16S rRNA sequences from that pool using PCR. Recall that PCR consists of repeated melting, annealing, and extension steps. During the annealing step of PCR, primers should largely bind to the target sequence, but some may bind to off-target sequences as well. Specificity of binding is controlled by both primer design and reaction conditions.
During the reaction itself, we can improve the reliability and accuracy of PCR by two key methods. The first is the use of a highly specific polymerase, one with either engineered or inherent hot start properties. Hot start means that the polymerase is inactive at low temperatures. Polymerases officially called "hot start" are negligibly active at room temperature, allowing reaction assembly without chilling, and also exhibit relatively lower activity at typical annealing temperatures (50-55 ° C) than other polymerases do. These properties tend to reduce binding of primers to non-target DNA. The polymerase you will use has much lower annealing temperature activity than the laboratory workhorse Taq, but is not officially a hot start polymerase. One important point to note here is that the target DNA (DNA of interest) may be present at a low concentration compared to the DNA in the sample as a whole, especially in complex mixtures, thus increasing opportunities for non-specific binding.
The second performance enhancer we will use is bovine serum albumin (BSA), which is especially important for amplifying DNA originating from a stool sample. Recall that stool samples are replete with inhibitors, including those that bind directly to DNA polymerases. As you saw if you clicked on the Kreader paper linked on Day 2, BSA itself binds many inhibitors of PCR, thus acting as a competitor. We would much rather that inhibitors bind the BSA than bind the polymerase and interfere with its function! BSA is hydrophobic and somewhat positively charged, making it a great non-specific binder of proteins that we will use time and again in 20.109.
Next time, you will visualize your entire reaction mixture in a gel, and if need be excise and purify the band of the correct size (~1400 bp) to isolate it from any non-specific products that occur in spite of the above precautions. Gel electrophoresis is a technique used to separate large molecules by size using an applied electrical field and appropriate sieving matrix. DNA fragments are typically separated in gels composed of agarose, a seaweed-derived polymer (see figure, below left). To prepare these gels, molten agarose is poured into a horizontal casting tray containing a comb. Once the agarose has solidified, the comb is removed, leaving wells into which the DNA sample can be loaded. The loaded DNA samples are then pulled through the matrix when a current is applied across it. Specifically, DNA molecules are negatively charged due to their phosphate backbones, and thus travel toward the positive charge at the far end of the gel (see figure, below right).
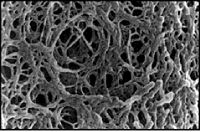
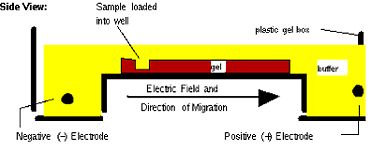
Although all DNA molecules travel in the same direction during gel electrophoresis, they do so at different rates: larger molecules get entwined in the matrix and retarded, while smaller molecules wind through the matrix more quickly and thus travel further from the well. Ultimately, fragments of similar length accumulate into “bands” in the gel. Bands of DNA are usually visualized by adding the fluorescent dye ethidium bromide (or newer alternatives such as SYBR Safe) to agarose gels. This dye intercalates between the bases of DNA, allowing DNA fragments to be located in the gel under UV light and photographed. The intensity of the band reflects the concentration of molecules that size, although there are upper and lower limits to the sensitivity of dyes. Because of its interaction with DNA, ethidium bromide is a powerful mutagen and will interact with the DNA in your body just as it does with any DNA on a gel. You should always handle all gels and gel equipment with nitrile gloves. Agarose gels with ethidium bromide must be disposed of as hazardous waste.
One parameter that affects the way DNA travels through a gel is the pore size, which is in turn affected by both the weight percent of the gel and the type of agarose used. Because we are separating large DNA fragments (> 1 Kbp) in the bacteria experiment, a low-to-medium percentage (namely 1.5 %) gel is appropriate. In the microsporidia experiment, small fragments (~ 0.1 Kbp) are expected and thus a high percentage (namely 3%) gel will be used. Moreover, we will use high-resolution (HR) agarose; its low viscosity means that high weight percent solutions are tractable to work with, and that the solidified gel remains pliable rather than brittle. HR agarose can be prepared by chemically modifying and/or partially depolymerizing natural agarose (as described here).
The PCR will continue during the whole lab period. In the meantime, we will discuss a journal article, both to learn more about investigations of the microbiome and to become comfortable reading and discussing the primary scientific literature. You will also hear from our oral presentation instructor on how to give a good talk, and get immediate feedback on your informal presentation of one slide. In 2-3 weeks, you will each present an article on your own.
Protocols
Part 1: Prepare PCR to detect bacterial 16S
- Begin by carefully labeling each PCR tube that you will use with the date, sample name, and a unique symbol and/or color for quick identification. Filling in the cap tab with your team color will usually suffice.
- Pre-chill the tubes on a cold block.
- You and your partner can now prepare and share a so-called "master mix," which contains every PCR ingredient except the template and the polymerase. Prepare enough for the number of reactions you need to run, plus an additional 10%. In addition to the two DNA-containing reactions, you should prepare a no-template control that contains pure water without any plasmid. Feel free to use the table below for your calculations.
- When the master mix is not in use, keep it on ice.
- What do you expect to see in the no-template control case?
- Combine 45 μL of master mix, 5 μL of template, and 1 μL of PfuUltra polymerase in a PCR tube. When everyone's reactions are ready, they will undergo the cycling conditions listed below.
- Add the master mix first, because the template alone may freeze. Then add template and polymerase, and finally (gently!) mix the reaction with a larger pipet.
| Reagent | Amount for 1 reaction (μL) | Amount for 3 reactions + 10% |
|---|---|---|
| PfuUltra buffer (10X stock) | 5 | |
| Primer mix | 1 | |
| dNTPs | 1 | |
| 5% BSA (100X stock) | 0.5 | |
| Water | 37.5 | |
| DNA template | 5 | N/A |
| Segment | Cycles | Temperature (° C) | Time |
|---|---|---|---|
| 1 | 1 | 95 | 5 min |
| 2-4 | 35 | 95 | 1 min |
| 51 | 1 min | ||
| 72 | 2 min | ||
| 5 | 1 | 72 | 10 min |
| 6 | 1 | 4 | indefinite |
Part 2: WAC Session
Today you will hear a lecture on preparing your journal club presentations.
Part 3: Journal article discussion
Scientific papers are dense and often time-consuming to read and understand, but with practice, you will find strategies that improve your comprehension efficiency. Here's one tip to get you started: when reading newly reported results, be sure to refer to the associated figures frequently, because visual information is often easier to take in than purely verbal descriptions.
Technical Background
Discussion Topics
Writing
As you read the paper by Koenig et al., consider not only its scientific content, but also the authors' writing style (perhaps not all on one read!). Sketch out answers to the questions below (right on the paper if you wish). Your answers will not be collected, but you may be called on in discussion to share your ideas.
- What functional elements does the abstract contain? As a whole, did the abstract make you want to read the paper?
- Now consider the Introduction section.
- What is the topic and/or function of each paragraph?
- How closely does this introduction conform to the suggested three-section structure described in the class scientific writing guidelines (LINK)? Is there too much or too little information or emphasis on any particular topic?
- What purpose(s) do the citations serve?
- Now consider the Results section.
- What purpose do the sub-section titles serve? Which ones do so most effectively?
- Can you find one or more examples of paragraphs with effective introductory and concluding sentences, according to the description here?
- How about examples with ineffective opening and closing sentences? How might you improve these?
- Are there any parts in the Results that you think belong in the Discussion instead, according to the descriptions here and here ?
- Finally, consider the Discussion section.
- What is the topic and/or function of each paragraph?
- Is there too much or too little information or emphasis on any particular topic?
- What purpose(s) do the citations serve?
Content
You were previously assigned one of the topics below to present to and discuss with the rest of the class. We'll now break to listen to Atissa's talk about giving talks, then give you some time to revise the slide that you prepared, and finally go through the slides and topics one by one. Per slide, we'll first discuss scientific content, and then give you feedback about your slide design and presentation -- briefly and informally.
- Overview and wrap-up? Supplementary figures?
- Figure 1
- Figure 2
- Figure 3
- Figure 4
- Figure 5
- Figure 6A (+S4?)
- Figure 6B (+S4?)
For next time
Reagent list
- PfuUltra polymerase and buffer from Agilent
- Primers
- intermediate stock concentration is 5 μM for each primer (5 μM total), diluted from individual 100 μM long-term stocks
- F8-27 sequence: 5' AGAGTTTGATCCTGGCTCAG
- R1392-1407 sequence: 5' ACGGGCGGTGTGTACA
