SBB12Ntbk-Robert D O'Dowd: Difference between revisions
| (18 intermediate revisions by the same user not shown) | |||
| Line 1: | Line 1: | ||
__NOTOC__ | __NOTOC__ | ||
==~~!~~== | ==~~!~~== | ||
{{SBB12_part_sbb1231}} <br> {{SBB12_part_sbb1214}} | |||
==[[User:Robert D O'Dowd|Robert D O'Dowd]] 2/16== | ==[[User:Robert D O'Dowd|Robert D O'Dowd]] 2/16== | ||
sbb1214 | '''sbb1214''' | ||
PCA on o16, o17, o12. Followed PCA protocol[http://http://openwetware.org/wiki/Template:SBB-PCA] | PCA on o16, o17, o12. Followed PCA protocol[http://http://openwetware.org/wiki/Template:SBB-PCA] | ||
| Line 9: | Line 11: | ||
Created 100uM dilutions of o16 o17 o12. | Created 100uM dilutions of o16 o17 o12. | ||
nmol weight x10 = uL of ddH2O need to add to get to 100 mM. | |||
Label tube 1214. Place on PCA rack. | Label tube 1214. Place on PCA rack. | ||
sbb1231 | '''sbb1231''' | ||
PCR on ca998/rdoSOECI_TM-R and pBca 9145-jtk2768 Label CI TM 1231 | PCR on ca998/rdoSOECI_TM-R and pBca 9145-jtk2768 Label CI TM 1231 | ||
| Line 27: | Line 29: | ||
==[[User:Robert D O'Dowd|Robert D O'Dowd]] 2/17== | ==[[User:Robert D O'Dowd|Robert D O'Dowd]] 2/17== | ||
sbb1214 | '''sbb1214''' | ||
Zymo cleanup-small fragment cleanup. [http://openwetware.org/wiki/Template:SBB-Protocols_Zymo2] Label 1214 PCA1 | Zymo cleanup-small fragment cleanup. [http://openwetware.org/wiki/Template:SBB-Protocols_Zymo2] Label 1214 PCA1 | ||
| Line 37: | Line 39: | ||
Could not conduct SOEPCR gel or setup. Doing next tuesday. | Could not conduct SOEPCR gel or setup. Doing next tuesday. | ||
sbb1231 | '''sbb1231''' | ||
Did not run any experiments | Did not run any experiments | ||
| Line 43: | Line 45: | ||
==[[User:Robert D O'Dowd|Robert D O'Dowd]] 2/21== | ==[[User:Robert D O'Dowd|Robert D O'Dowd]] 2/21== | ||
sbb1231 | '''sbb1231''' | ||
Prep 6 uL product w/ 4 uL loading buffer Placed on wells 9, 10 labeled ROTP and ROCT | Prep 6 uL product w/ 4 uL loading buffer Placed on wells 9, 10 labeled ROTP and ROCT | ||
Picture taken of gel->does not look correct | Picture taken of gel->does not look correct | ||
Lane 9 has W shaped smear | Lane 9 has W shaped smear | ||
I do not believe my recordings. Either I switched what lanes I believed was which or one of my products failed. I will switch assumptions and assum 9 ROCT and 10 ROTP | I do not believe my recordings. Either I switched what lanes I believed was which or one of my products failed. I will switch assumptions and assum 9 ROCT and 10 ROTP | ||
Cut out band which correspond to correct size. | Cut out band which correspond to correct size. | ||
Cut out extra band. 1K band maybe from lane 9 is ROCT | Cut out extra band. 1K band maybe from lane 9 is ROCT | ||
Possible that 1K hidden in smear is actual TP product | Possible that 1K hidden in smear is actual TP product | ||
No 500 band appeared in lane 10 so its doubtful. | No 500 band appeared in lane 10 so its doubtful. | ||
Zymo gel purification | Zymo gel purification | ||
label 1231ROTP, 1231ROCT, 1231MaybeTP | label 1231ROTP, 1231ROCT, 1231MaybeTP | ||
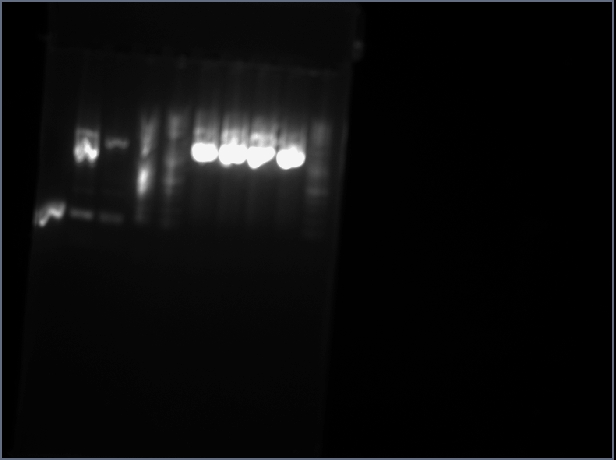
[[Image:NEB_2-log_ladder.gif]] | [[Image:NEB_2-log_ladder.gif]][[Image:2012_02_21_gel1_ssb2012spring.jpg]]<br> | ||
[[Image:2012_02_21_gel1_ssb2012spring.jpg]]<br> | |||
Lanes:<br> | Lanes:<br> | ||
| Line 77: | Line 85: | ||
|} | |} | ||
sbb1214 | '''sbb1214''' | ||
Cloning step. Skip reassembly. Product named 1214 Cloning | |||
Followed digestion of wobble product protocol [http://openwetware.org/wiki/Template:SBB-Protocols_Enz1] except digested with | |||
NheI/BamHI | |||
Place in incubator at 1120 | Place in incubator at 1120 | ||
Store 1214 Cloning | Store 1214 Cloning | ||
==[[User:Robert D O'Dowd|Robert D O'Dowd]] 2/23== | ==[[User:Robert D O'Dowd|Robert D O'Dowd]] 2/23== | ||
sbb1231 | '''sbb1231''' | ||
SOE PCR Round 3 | SOE PCR Round 3 | ||
PCR 1231ROCT + 1231ROTP with ca998/g00101 | PCR 1231ROCT + 1231ROTP with ca998/g00101 | ||
Result labeled 1231PCR3 | Result labeled 1231PCR3 | ||
Placed in PCR rack for 1K-2K | Placed in PCR rack for 1K-2K | ||
sbb1214 | '''sbb1214''' | ||
Taking digested product from tuesday->zymo column [http://openwetware.org/wiki/Template:SBB-Protocols_Zymo2] | Taking digested product from tuesday->zymo column [http://openwetware.org/wiki/Template:SBB-Protocols_Zymo2] | ||
Too much liquid for 1 column. Performing 2 rounds. | Too much liquid for 1 column. Performing 2 rounds. | ||
Added 50 uL H2O. This amount was not correct should have only eluded w/8uL. Possible that product is too diluted for later steps to work. | Added 50 uL H2O. This amount was not correct should have only eluded w/8uL. Possible that product is too diluted for later steps to work. | ||
Ligate products | Ligate products | ||
Incubate on benchtop for 30 min 11:12-11:42 | Incubate on benchtop for 30 min 11:12-11:42 | ||
Waiting until Friday for transformation | Waiting until Friday for transformation | ||
If there is an issue, it will likely be due to 6x dilution of vector | If there is an issue, it will likely be due to 6x dilution of vector | ||
Can restart at 1214Cloning | Can restart at 1214Cloning | ||
==[[User:Robert D O'Dowd|Robert D O'Dowd]] 2/24== | ==[[User:Robert D O'Dowd|Robert D O'Dowd]] 2/24== | ||
sbb1231 | '''sbb1231''' | ||
Run gel for PCR product | Run gel for PCR product | ||
Gel1 Lane 10 Labeled ROSOE3 1231 | Gel1 Lane 10 Labeled ROSOE3 1231 | ||
Place 1231 PCR3 back into box of products | Place 1231 PCR3 back into box of products | ||
| Line 132: | Line 156: | ||
Cut out band at 2K place in tube labeled 1231SOE3 600uL ADB. | Cut out band at 2K place in tube labeled 1231SOE3 600uL ADB. | ||
Melt and place on ice | Melt and place on ice | ||
2 bands show up. one considerably brighter and heavier, Selected smaller band because it looks closer to the target 1800 bp I expect. | 2 bands show up. one considerably brighter and heavier, Selected smaller band because it looks closer to the target 1800 bp I expect. | ||
sbb1214 | '''sbb1214''' | ||
Need to redo ligation | Need to redo ligation | ||
Incubating on benchtop for 30 min 12:30-1:00 | Incubating on benchtop for 30 min 12:30-1:00 | ||
Label product 1213lig | Label product 1213lig | ||
place on ice after 30 min | place on ice after 30 min | ||
mix 30 uL KCM | mix 30 uL KCM | ||
Transfer 70 uL cell mixture to ligation tube. Wait 10 min 117-127 | Transfer 70 uL cell mixture to ligation tube. Wait 10 min 117-127 | ||
Probably dont have enough time to finish plating | Probably dont have enough time to finish plating | ||
| Line 149: | Line 181: | ||
==[[User:Robert D O'Dowd|Robert D O'Dowd]] 2/28== | ==[[User:Robert D O'Dowd|Robert D O'Dowd]] 2/28== | ||
sbb1231 | '''sbb1231''' | ||
Complete zymo cleanup from product on Friday | Complete zymo cleanup from product on Friday | ||
Label purified product 1231SOEPur | Label purified product 1231SOEPur | ||
Set up digestion from 1231SOEPur. Label 1231 dig. place at 37 at 1040-1140 | Set up digestion from 1231SOEPur. Label 1231 dig. place at 37 at 1040-1140 | ||
Set up gel. 1202-1232 | Set up gel. 1202-1232 | ||
I expect to see a band at 1890 | I expect to see a band at 1890 | ||
| Line 178: | Line 214: | ||
Dont see anything. Need to redo PCRs | Dont see anything. Need to redo PCRs | ||
sbb1214 | '''sbb1214''' | ||
Redo ligation for sbb1214 | Redo ligation for sbb1214 | ||
Label 1214 Lig. 1028-1058 bench top incubate. | Label 1214 Lig. 1028-1058 bench top incubate. | ||
Thaw cells and label cell prep. add 50 uL H2O. | Thaw cells and label cell prep. add 50 uL H2O. | ||
Transfer to new tube labeled 1214 Trans. Place on ice 1111-1121 | Transfer to new tube labeled 1214 Trans. Place on ice 1111-1121 | ||
Heatshok at 42C for 90 sec | Heatshok at 42C for 90 sec | ||
Place in shaker 1127-1227 | Place in shaker 1127-1227 | ||
Cutting incubatinon time by 15 min in order to complete during class period. 1212 | Cutting incubatinon time by 15 min in order to complete during class period. 1212 | ||
Cells plated on Spec plates using alcohol sterilized wand. | Cells plated on Spec plates using alcohol sterilized wand. | ||
Plate labeled 2/28 RDO1214 pBca1825_1214 | Plate labeled 2/28 RDO1214 pBca1825_1214 | ||
Plate given to Zach | Plate given to Zach | ||
==[[User:Robert D O'Dowd|Robert D O'Dowd]] 3/1== | ==[[User:Robert D O'Dowd|Robert D O'Dowd]] 3/1== | ||
sbb1231 | '''sbb1231''' | ||
Run gel on PCR products CI_TM and TM_PhoA and make sure bands show up | Run gel on PCR products CI_TM and TM_PhoA and make sure bands show up | ||
Redo SOEPCR | Redo SOEPCR | ||
Run gel. Lane 8 TMPhoA Lane 9 CITM | Run gel. Lane 8 TMPhoA Lane 9 CITM | ||
| Line 220: | Line 267: | ||
Did not turn out well. Need to redo PCR | Did not turn out well. Need to redo PCR | ||
sbb1214 | '''sbb1214''' | ||
One sample vial of cultured cells found 1214-1. Apparently only 1 colony was picked. | One sample vial of cultured cells found 1214-1. Apparently only 1 colony was picked. | ||
Followed DNA Miniprep procedure. [http://openwetware.org/wiki/Template:SBB-Protocols_Micro3] Label all tubes used in process 1213_1RO | Followed DNA Miniprep procedure. [http://openwetware.org/wiki/Template:SBB-Protocols_Micro3] Label all tubes used in process 1213_1RO | ||
Final product labeled 1213-1RO mPrep | Final product labeled 1213-1RO mPrep | ||
Need to run an analytical gel next time | Need to run an analytical gel next time | ||
==[[User:Robert D O'Dowd|Robert D O'Dowd]] 3/2== | ==[[User:Robert D O'Dowd|Robert D O'Dowd]] 3/2== | ||
'''sbb1231''' | |||
Because final expected product did not show up in gel last time, I need to start over. Reconstructing CITM and TM PhoA. | |||
PCR ca998/RDOSOECITM_R on pBca9145jtk2768. Mixture placed on 500-1k Rack | |||
PCR RdoSOETMPhoA_F/g00101 on pBjh1601CK Bjh2128. Mixture placed on 1k-2k rack | |||
'''sbb1214''' | |||
Since it is recommended to have 2 clones and I only 1 colony was picked, I should run another transformation in order to get more clones | |||
Digest pBca9525 Bca 1834 Nhe1 BamH1 and ligate with 1214dig | |||
Not enough time to transform and plate cells. | |||
Running an analytical gel with EcoRI XholI. Digest placed in incubator 103-133 | |||
[[Image:NEB_2-log_ladder.gif]][[Image:2012_03_02_gel2_ssb2012spring.jpg]]<br> | |||
Lanes:<br> | |||
{| border="1" cellpadding="2" | |||
!width="1"|1 | |||
!width="1"|2 | |||
!width="1"|3 | |||
!width="1"|4 | |||
!width="1"|5 | |||
!width="1"|6 | |||
!width="1"|7 | |||
!width="1"|8 | |||
!width="1"|9 | |||
!width="1"|10 | |||
|- | |||
||AJ A+B||AJ A+B||MM 1232-1||MM 1232-2||MM 1215-2||RO MP 1214||Ladder||MDP2||MDP1||MDL2 | |||
|} | |||
==[[User:Robert D O'Dowd|Robert D O'Dowd]] 3/6== | |||
'''sbb1231''' | |||
Need to run gel to retrieve and purify SOEPCR products | |||
Lanes 1 (CITM) and 2 (TMPhoA) on Gel 3 | |||
[[Image:NEB_2-log_ladder.gif]][[Image:2012_03_05_gel3_ssb2012spring.jpg]]<br> | |||
Perform Zymo gel purification | |||
1231 CITM2 and 1231 TMPhoA2 stored | |||
Solution labeled 1231 SOE3. Placed on 1-2K rack | |||
'''sbb1214''' | |||
Looking back on my notes, I did not run the second round of PCA w/ o16 and o12 on pca1. | |||
Restarting | |||
PCA1 with o12, o16, o17 | |||
Note from 3/15 | |||
My results from the analytical gel would seem to indicate that I had performed the correct reactions even though I did not record them in my notebook. I continue with the restart anyways. | |||
==[[User:Robert D O'Dowd|Robert D O'Dowd]] 3/8== | |||
'''sbb1231''' | |||
1231 SOE3 digested with EcoRI and BamHI. Solution labeled 1231 Dig | |||
Placed in thermocycler 1007-1107 | |||
There is not enough time to ligate and plate. | |||
Next week plan: | |||
Digest 1h -> Gel 30m -> Zymo 15m -> Ligate 30m -> Transform 1h -> Plate 15m | |||
'''sbb1214''' | |||
Set up amplification for Pca1 mixture | |||
Label 1214Pca1 amp and place on Pca2 Rack for next stage | |||
Next week plan: | |||
Zymo -> Digestion -> Ligation -> Transform -> Plating | |||
Can run steps in parallel with 1231 at the ligation stage | |||
==[[User:Robert D O'Dowd|Robert D O'Dowd]] 3/13== | |||
'''sbb1231''' | |||
1231 SOE3 digested with EcoRI and BamHI to get 1231 Dig | |||
Placed in thermocycler from 1022-1122 | |||
Run gel. Lane 2 (RO 1231 dig) | |||
Save image | |||
Cut out the bright band in Lane 2 | |||
Add 600 mL ADB buffer and store for next time | |||
'''sbb1214''' | |||
1214Pca1 amp purified using zymo small fragment procedure. | |||
Labeled 1214 pur | |||
Begin digestion with NheI and BamHI. Result is 1214 dig. | |||
Placed in thermocycler 1022-1122 | |||
1214 dig run on gel. Lane 1 (RO 1214 dig) | |||
Add 600 mL ADB and store for next time | |||
[[Image:NEB_2-log_ladder.gif]][[Image:2012_03_13_gel3_ssb2012spring.jpg]]<br> | |||
Lanes:<br> | |||
{| border="1" cellpadding="2" | |||
!width="1"|1 | |||
!width="1"|2 | |||
!width="1"|3 | |||
!width="1"|4 | |||
!width="1"|5 | |||
!width="1"|6 | |||
!width="1"|7 | |||
!width="1"|8 | |||
!width="1"|9 | |||
!width="1"|10 | |||
|- | |||
||RO 1214 Dig||RO 1231 Dig||AJ A+B PCR digest||****||Lader||KA||KA||KA||KA||ladder | |||
|} | |||
==[[User:Robert D O'Dowd|Robert D O'Dowd]] 3/15== | |||
'''sbb1231''' | |||
Zymo gel cleanup on RO 1231 Dig | |||
Run ligation reaction. | |||
pBca9525-Bca1834 digested with BamHI and EcoRI and ligated with RO 1231 Dig. Product is RO 1231 Lig | |||
Place on bench for 30 min 1031-1101 | |||
Add KCM to competent cells. Place on Ice for 10 min 1110-1120 | |||
Transform cells with RO 1231 Lig in thermomcycler at 42 C for 90 seconds. | |||
Place on ice for 1 min | |||
Place in incubator for 1 hour 1126-1226 | |||
Spilled my transformed cells and lost about half. This may cause contamination or not enough cells being plated | |||
Plate on Spec plates and label RDO 1231 3/15 | |||
'''sbb1214''' | |||
Add 250 mL isopropanol. Proceed with Zymo small frag gel cleanup. Label resulting solution RO 1214 Dig | |||
Digest pBca9525-Bca1834 with NheI and BamHI. Ligate this with RO 1214 Dig. Result is RO 1214 Lig | |||
Incubate on bench for 30 in. 1031-1101. | |||
Add cells to ligation. Let it sit on ice 10 min 1110-1120 | |||
Place in thermocycler at 42 C for 90 seconds | |||
Place back on ice for 1 min | |||
Place in incubator for 1 hour. 1126-1226 | |||
Plate cells on Spec plates and label them RDO 1214 3/15 | |||
==[[User:Robert D O'Dowd|Robert D O'Dowd]] 3/20== | |||
'''sbb1231''' | |||
4 Colonies were picked by Zach. | |||
Turns out none of the cell cultures were viable and the plates I created were totally unusable. There are 2 large red clusters where cells survived. The colonies that were picked were likely contamination from when I leaked. It is also possible that the transformation totally failed. | |||
Experiment stopped. sbb1231 inconclusive. | |||
'''sbb1214''' | |||
4 colonies picked. All 4 cultures are viable. | |||
Performing miniprep on all 4. [http://openwetware.org/wiki/Template:SBB-Protocols_Micro3] | |||
Resulting solutions were labeled RO 1214 1, RO 1214 2, RO 1214 3, RO 1214 4 | |||
Running analytical digests on miniprepped plasmids | |||
RO 1214 1 and 2 | |||
Digest with XbaI and NheI | |||
Expect bands at 3161 and 456 if successful and 3161 and 1174 if unsuccessful | |||
RO 1214 3 and 4 | |||
Digest with BamHI and EcoRI | |||
Expect bands at 2472 and 1145 if successful and 2472 and 1863 if unsuccessful | |||
Not enough time to incubate and run the gel. | |||
Performing the gel on Thursday. | |||
Removing 1231ROCT, 1231PCR3, 1231CITM, 1214Pca1, 1231TMPhoA from storage in order to make space. | |||
==[[User:Robert D O'Dowd|Robert D O'Dowd]] 3/22== | |||
'''sbb1231''' | |||
Not enough time to complete the experiment. I can redo the transformation and plating to at least figure out if my products were viable. If white colonies appear the Friday on a plate, I can say that the transformation at least happened. | |||
Rerun ligation 1031-1101. pBca9525-Bca1834 digested with BamHI and EcoRI and ligated with RO 1231 Dig. Product is RO 1231 Lig2. | |||
Competent cells thawed and prepared. Mixed with RO 1231 Lig2. Product is 1231 trans. Placed on ice for 10 min. 1113-1123 | |||
Incubate 1127-1127. | |||
Cutting incubation 7 minutes early due to time. | |||
Cells plated on Spec plates. Labeled RDO1231 3/22 No need to pick | |||
'''sbb1214''' | |||
Mapping miniprep | |||
Thermocycler 30 min 1016-1046 | |||
Not sure if 1 and 2 will work. The XbaI enzyme solution is almost empty and less than .5 uL was added. | |||
Run gel on lanes 1,2,3,4. | |||
[[Image:NEB_2-log_ladder.gif]][[Image:2012_03_22_gel1_ssb2012spring.jpg]]<br> | |||
Lanes:<br> | |||
{| border="1" cellpadding="2" | |||
!width="1"|1 | |||
!width="1"|2 | |||
!width="1"|3 | |||
!width="1"|4 | |||
!width="1"|5 | |||
!width="1"|6 | |||
!width="1"|7 | |||
!width="1"|8 | |||
!width="1"|9 | |||
!width="1"|10 | |||
|- | |||
||none||RO 1214 1||RO 1214 2||RO 1214 3||RO 1214 4||2-log||AJ digest||DC A+B||JW A10||none | |||
|} | |||
All samples result in bands in the expected locations. Submitting 1214 2 and 1214 4 for sequencing. | |||
Latest revision as of 18:39, 12 April 2012
~~!~~
Partname: sbb1231 Featurename: cI-TM Genename: cI, other Source: Lambda phage / other
This one is a little different. You aren't putting a homodimer onto ToxR; you're building out a different homodimerization system. In this experiment, you are trying to reproduce some experiments based on PMID 9671551. In their study, they append different dimerizing transmembrane (TM) segments to the lambda repressor DNA binding domain fragment (cI') and see whether CI's activity is restored. In one case, they discuss making fusion proteins such as:
Nterm-cI'-linker-TM-PhoA-Cterm
That would be a nice construct to have because you can do 2 assays with it: transcriptional repression from cI-dependent promoters, and periplasmic targetting with alkaline phosphatase.
Your design of this part depends on your reading of the paper and the availability of materials from which you could construct the things they describe. You'll probably want to discuss the design of your part with JCA after you have read and made sense of the paper. Your final construct should be a normal BglBricks part in plasmid pBca9525. You most likely will want to piece the part together from two (or more) pcr products. The cI construct can be amplified from pBca9145-jtk2768. You can get PhoA from plasmid pBjh1601CK-Bjh2128.
Construction File
PCR ca998/rdoSOECI_TM-R on pBca9145-jtk2768 (cifrag, 583bp) PCR rdoSOETM_PhoA-F/g00101 on pBjh1601CK-Bjh2128 (phofrag, 1327bp) PCR ca998/g00101 on cifrag+phofrag (pcrpdt_sbb1231, 1890bp) Digest pBca9525-Bca1834 (EcoRI/BamHI, L, vectdig) Digest pcrpdt_sbb1231 (EcoRI/BamHI, L, pcrdig) Ligate pcrdig and vectdig, product is pBca9525-sbb1231 ---- >rdoSOECI_TM-R Combining CI'head and MalF TM CCAACCAGCAGACCCAGCAGACCCAGAACAGACCATTTACGTTTAGATCCgcttacccatctctccgcatc >rdoSOETM_PhoA-F Combining MalF TM with PhoA CTGCTGGGTCTGCTGGTTGGTTACCTGGTTGTTCTGATGTACGGATCTtctagaATTGATGCCCTGCCGCTGAC >pcrpdt GTATCACGAGGCAGAATTTCAGATAAAAAAAATCCTTAGCTTTCGCTAAGGATGATTTCTGGAATTCATGAGATCTGTTAATGCTAGCTCAGTCCTAGGGACTCTGCTAGCGGATCTAGACATAAAAACGGCAAAGTATGAGCACAAAAAAGAAACCATTAACACAAGAGCAGCTTGAGGACGCACGTCGCCTTAAAGCAATTTATGAAAAAAAGAAAAATGAACTTGGCTTATCCCAGGAATCTGTCGCAGACAAGATGGGGATGGGGCAGTCAGGCGTTGGTGCTTTATTTAATGGCATCAATGCATTAAATGCTTATAACGCCGCATTGCTTGCAAAAATTCTCAAAGTTAGCGTTGAAGAATTTAGCCCTTCAATCGCCAGAGAAATCTACGAGATGTATGAAGCGGTTAGTATGCAGCCGTCACTTAGAAGTGAGTATGAGTACCCTGTTTTTTCTCATGTTCAGGCAGGGATGTTCTCACCTGAGCTTAGAACCTTTACCAAAGGTGATGCGGAGAGATGGGTAAGCGGATCTAAACGTAAATGGTCTGTTCTGGGTCTGCTGGGTCTGCTGGTTGGTTACCTGGTTGTTCTGATGTACGGATCTTCTAGAATTGATGCCCTGCCGCTGACCGGACAATATACCCACTATGCGCTGAATAAAAAAACCGGCAAACCAGATTACGTGACCGATTCCGCCGCCTCAGCCACCGCGTGGTCCACAGGGGTGAAAACCTACAACGGCGCGCTGGGGGTCGATATTCATGAAAAAGATCATCTGACCATTCTCGAAATGGCGAAGGCCGCAGGTCTGGCTACCGGCAACGTCTCGACCGCGGAGTTGCAGGACGCCACCCCGGCAGCGCTGATTGCCCATGTCACCTCGCGCAAATGCTATGGCCCGACGGCGACCAGCGAAAAATGTCCGACAAACGCGCTGGAAAAGGGCGGCAAAGGCTCGATTACTGAACAGCTCCTGAACGCGCGGGCGGACGTCACCCTGGGCGGCGGCGCGAAAACCTTCGCCGAGACGGCAACCGCCGGCGAGTGGCAGGGGAAAACGCTGCGTGAGCAGGCGCAGATGCGCGGCTACCAGTTAGTCGGCGATGCCGCCTCTTTAGCGGCGATCGACGAAGCGAATCAGGACAAACCGCTGCTGGGACTGTTCGCAGAGGGGAATATGCCGGTGCGCTGGGAAGGGCCGAAGGCCTCGTACCACGGCAATATTGATAAACCTGCCGTGACCTGTACGCCAAATCCGAAACGCAATGACGCGATTCCCACGCTGGCGCAGATGACCGACAAAGCCATCGAACTGTTGAGCAAAAATGAGCGGGGCTTCTTTTTACAGGTGGAAGGCGCGTCTATCGATAAGCAGGATCACGCCGCGAACCCGTGCGGACAGATTGGCGAGACGGTGGATCTTGATGAAGCCGTACAGAGGGCGCTGGCCTTTGCGAAGAAAGACGGCAATACGCTGGTGATCGTTACCGCCGATCATGCTCATGCCAGCCAGATCGTGGCGCCCGAAACGAAAGCGCCGGGGCTGACGCAGGCGCTGAACACCAAAGATGGCGCGGTGATGGTCATCAGCTACGGCAACTCGGAAGAAGATTCCCAGGAGCATACCGGCAGCCAGCTGCGCATCGCCGCTTACGGGCCGCATGCGGCGAACGTGGTCGGGCTGACCGATCAAACCGATCTGTTCTACACCATGAAAGCGGCGCTGGGCCTGAAGTAATGGATCTAAAAAAAAACCCCGCCCCTGACAGGGCGGGGTTTTTTTTAGGATCCTAACTCGACGTGCAGGCTTCCTCGCTCACTGACTCGCTGCGCTCGGTCGTTCGGCTGCGGCGAGCGGTATCAGCTCACTCAAAGGCGGTAAT
Partname: sbb1214 Featurename: lz_AAAA-3 Genename: leucine zipper variant Source: Synthetic, see PMID:12459719
This part encodes a leucine zipper
We will be using a set of leucine zippers as homodimer domains. In total, we'll have 8 of them all with different Kd's for homodimerization. However, the sequences only differ at the same 4 amino acid sites. This will allow us to test whether the activation of ToxR is dependent on the Kd of the homodimer that is attached to it. These sequences are entirely synthetic, but all should encode the following peptide:
VKELEDKNEELLS XX YH XX NEVARLKKLVGERGGC*
Where the x's are the amino acids I tell you. So, if I gave you the lz_IILK zipper to make, the peptide you should encode is: VKELEDKNEELLSIIYHLKNEVARLKKLVGERGGC*
Here is an example of what your trying to make pBca9525-sbb1230.
This part encodes a homodimer domain
You are going to put the homodimer on the C-terminus of a truncated ToxR. For this part, you're going to go off BioBricks. Kindov. Your final product will be a BioBrick, but you are going to use an ad hoc procedure to create it. You should examine this illustration which refers to two sequences: pBca9525-Bca1834 and pBca9525-Bca1839. The important thing here is that you think through the translation frame of the final product, and make sure that you add some linker between your sequence and the sequence 5' to it. You should also remove the start methionine from you homodimer, unless your project description explicitly says not to. You should include the stop codon.
The product of your construction file should resemble pBca9525-Bca1839 in terms of it encoding a full expression cassette containing a ToxR fusion protein with your designed feature. If you were given part sbb1293, your product would be called pBca9525-sbb1293.
Construction File
PCA1 on o16,o17,o12 (pca1) PCA2 with o16/o12 on pca1 (139 bp, pca2) Digest pca2 (NheI/BamHI, L, 1214dig) Digest pBca9525-Bca1834 (NheI/BamHI, L, vectdig) Ligate 1214dig + vectdig, product is pBca9525-sbb1214 ---- >o16 CCATAgctagcGGCAGTGGATCTGTTAAAGAAGCGGAAGACAAAGCAGAAGAAGCGCTGAGT >o17 CAAAGCAGAAGAAGCGCTGAGTATCATCTACCACCTGAAAAACGAAGTTGCTCGTCTGA >o12 CAGTAGGATCCTTAGCCGCCACGTTCGCCAACCAGTTTTTTCAGACGAGCAACTTCGTT >pca2 CCATAgctagcGGCAGTGGATCTGTTAAAGAAGCGGAAGACAAAGCAGAAGAAGCGCTGAGTATCATCTACCACCTGAAAAACGAAGTTGCTCGTCTGAAAAAACTGGTTGGCGAACGTGGCGGCTAAGGATCCTACTG
Robert D O'Dowd 2/16
sbb1214
PCA on o16, o17, o12. Followed PCA protocol[1]
Created 100uM dilutions of o16 o17 o12.
nmol weight x10 = uL of ddH2O need to add to get to 100 mM.
Label tube 1214. Place on PCA rack.
sbb1231
PCR on ca998/rdoSOECI_TM-R and pBca 9145-jtk2768 Label CI TM 1231
PCR on rdoSOETM_PhoA-F/g00101 and pBjh 1601CK-Bjh2128 Label TM PhoA 1231
Followed PCR protocol [2]
Place CI TM 1231 on 500-1K bp rack
Place Label TM PhoA 1231 on 1K-2K bp rack
Robert D O'Dowd 2/17
sbb1214
Zymo cleanup-small fragment cleanup. [3] Label 1214 PCA1
Amplification/phusion step. Template DNA added last. [4]
Template material and fragments for SOE placed in Eluded Products box.
Could not conduct SOEPCR gel or setup. Doing next tuesday.
sbb1231
Did not run any experiments
Robert D O'Dowd 2/21
sbb1231
Prep 6 uL product w/ 4 uL loading buffer Placed on wells 9, 10 labeled ROTP and ROCT
Picture taken of gel->does not look correct
Lane 9 has W shaped smear
I do not believe my recordings. Either I switched what lanes I believed was which or one of my products failed. I will switch assumptions and assum 9 ROCT and 10 ROTP
Cut out band which correspond to correct size.
Cut out extra band. 1K band maybe from lane 9 is ROCT
Possible that 1K hidden in smear is actual TP product
No 500 band appeared in lane 10 so its doubtful.
Zymo gel purification
label 1231ROTP, 1231ROCT, 1231MaybeTP
Lanes:
| 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 |
|---|---|---|---|---|---|---|---|---|---|
| Au PCA2 | TN sbb1213 | DC A | DC B | 2-log ladder | JS | MM | JW | RO TP | RO CT |
sbb1214
Cloning step. Skip reassembly. Product named 1214 Cloning
Followed digestion of wobble product protocol [5] except digested with NheI/BamHI
Place in incubator at 1120
Store 1214 Cloning
Robert D O'Dowd 2/23
sbb1231
SOE PCR Round 3
PCR 1231ROCT + 1231ROTP with ca998/g00101
Result labeled 1231PCR3
Placed in PCR rack for 1K-2K
sbb1214
Taking digested product from tuesday->zymo column [6]
Too much liquid for 1 column. Performing 2 rounds.
Added 50 uL H2O. This amount was not correct should have only eluded w/8uL. Possible that product is too diluted for later steps to work.
Ligate products
Incubate on benchtop for 30 min 11:12-11:42
Waiting until Friday for transformation
If there is an issue, it will likely be due to 6x dilution of vector
Can restart at 1214Cloning
Robert D O'Dowd 2/24
sbb1231
Run gel for PCR product
Gel1 Lane 10 Labeled ROSOE3 1231
Place 1231 PCR3 back into box of products
Lanes:
| 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 |
|---|---|---|---|---|---|---|---|---|---|
| MH PCRPDT 1218 | MH PCRDIG 1218 | empty | DC SB 1221 | Ladder | empty | empty | empty | GF PCA | RO SOE3 1231 |
Cut out band at 2K place in tube labeled 1231SOE3 600uL ADB.
Melt and place on ice
2 bands show up. one considerably brighter and heavier, Selected smaller band because it looks closer to the target 1800 bp I expect.
sbb1214
Need to redo ligation
Incubating on benchtop for 30 min 12:30-1:00
Label product 1213lig
place on ice after 30 min
mix 30 uL KCM
Transfer 70 uL cell mixture to ligation tube. Wait 10 min 117-127
Probably dont have enough time to finish plating
Toss materials and retry tuesday
Robert D O'Dowd 2/28
sbb1231
Complete zymo cleanup from product on Friday
Label purified product 1231SOEPur
Set up digestion from 1231SOEPur. Label 1231 dig. place at 37 at 1040-1140
Set up gel. 1202-1232
I expect to see a band at 1890
Lanes:
| 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 |
|---|---|---|---|---|---|---|---|---|---|
| empty | AJ A+B pcrpdt | empty | RD 1231 dig | empty | 2-log ladder | empty | empty | empty | empty |
Dont see anything. Need to redo PCRs
sbb1214
Redo ligation for sbb1214
Label 1214 Lig. 1028-1058 bench top incubate.
Thaw cells and label cell prep. add 50 uL H2O.
Transfer to new tube labeled 1214 Trans. Place on ice 1111-1121
Heatshok at 42C for 90 sec
Place in shaker 1127-1227
Cutting incubatinon time by 15 min in order to complete during class period. 1212
Cells plated on Spec plates using alcohol sterilized wand.
Plate labeled 2/28 RDO1214 pBca1825_1214
Plate given to Zach
Robert D O'Dowd 3/1
sbb1231
Run gel on PCR products CI_TM and TM_PhoA and make sure bands show up
Redo SOEPCR
Run gel. Lane 8 TMPhoA Lane 9 CITM
Lanes:
| 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 |
|---|---|---|---|---|---|---|---|---|---|
| Karin 1206 | 1224(1) | 1224(2) | 1224(3) | 1203(1) | ladder | AJ | RO TM PhoI | RO CITM | Bad Load |
Did not turn out well. Need to redo PCR
sbb1214
One sample vial of cultured cells found 1214-1. Apparently only 1 colony was picked.
Followed DNA Miniprep procedure. [7] Label all tubes used in process 1213_1RO
Final product labeled 1213-1RO mPrep
Need to run an analytical gel next time
Robert D O'Dowd 3/2
sbb1231
Because final expected product did not show up in gel last time, I need to start over. Reconstructing CITM and TM PhoA.
PCR ca998/RDOSOECITM_R on pBca9145jtk2768. Mixture placed on 500-1k Rack
PCR RdoSOETMPhoA_F/g00101 on pBjh1601CK Bjh2128. Mixture placed on 1k-2k rack
sbb1214
Since it is recommended to have 2 clones and I only 1 colony was picked, I should run another transformation in order to get more clones
Digest pBca9525 Bca 1834 Nhe1 BamH1 and ligate with 1214dig
Not enough time to transform and plate cells.
Running an analytical gel with EcoRI XholI. Digest placed in incubator 103-133
Lanes:
| 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 |
|---|---|---|---|---|---|---|---|---|---|
| AJ A+B | AJ A+B | MM 1232-1 | MM 1232-2 | MM 1215-2 | RO MP 1214 | Ladder | MDP2 | MDP1 | MDL2 |
Robert D O'Dowd 3/6
sbb1231
Need to run gel to retrieve and purify SOEPCR products
Lanes 1 (CITM) and 2 (TMPhoA) on Gel 3
Perform Zymo gel purification
1231 CITM2 and 1231 TMPhoA2 stored
Solution labeled 1231 SOE3. Placed on 1-2K rack
sbb1214
Looking back on my notes, I did not run the second round of PCA w/ o16 and o12 on pca1.
Restarting
PCA1 with o12, o16, o17
Note from 3/15
My results from the analytical gel would seem to indicate that I had performed the correct reactions even though I did not record them in my notebook. I continue with the restart anyways.
Robert D O'Dowd 3/8
sbb1231
1231 SOE3 digested with EcoRI and BamHI. Solution labeled 1231 Dig
Placed in thermocycler 1007-1107
There is not enough time to ligate and plate.
Next week plan:
Digest 1h -> Gel 30m -> Zymo 15m -> Ligate 30m -> Transform 1h -> Plate 15m
sbb1214
Set up amplification for Pca1 mixture
Label 1214Pca1 amp and place on Pca2 Rack for next stage
Next week plan:
Zymo -> Digestion -> Ligation -> Transform -> Plating
Can run steps in parallel with 1231 at the ligation stage
Robert D O'Dowd 3/13
sbb1231
1231 SOE3 digested with EcoRI and BamHI to get 1231 Dig
Placed in thermocycler from 1022-1122
Run gel. Lane 2 (RO 1231 dig)
Save image
Cut out the bright band in Lane 2
Add 600 mL ADB buffer and store for next time
sbb1214
1214Pca1 amp purified using zymo small fragment procedure.
Labeled 1214 pur
Begin digestion with NheI and BamHI. Result is 1214 dig.
Placed in thermocycler 1022-1122
1214 dig run on gel. Lane 1 (RO 1214 dig)
Add 600 mL ADB and store for next time
Lanes:
| 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 |
|---|---|---|---|---|---|---|---|---|---|
| RO 1214 Dig | RO 1231 Dig | AJ A+B PCR digest | **** | Lader | KA | KA | KA | KA | ladder |
Robert D O'Dowd 3/15
sbb1231
Zymo gel cleanup on RO 1231 Dig
Run ligation reaction.
pBca9525-Bca1834 digested with BamHI and EcoRI and ligated with RO 1231 Dig. Product is RO 1231 Lig
Place on bench for 30 min 1031-1101
Add KCM to competent cells. Place on Ice for 10 min 1110-1120
Transform cells with RO 1231 Lig in thermomcycler at 42 C for 90 seconds.
Place on ice for 1 min
Place in incubator for 1 hour 1126-1226
Spilled my transformed cells and lost about half. This may cause contamination or not enough cells being plated
Plate on Spec plates and label RDO 1231 3/15
sbb1214
Add 250 mL isopropanol. Proceed with Zymo small frag gel cleanup. Label resulting solution RO 1214 Dig
Digest pBca9525-Bca1834 with NheI and BamHI. Ligate this with RO 1214 Dig. Result is RO 1214 Lig
Incubate on bench for 30 in. 1031-1101.
Add cells to ligation. Let it sit on ice 10 min 1110-1120
Place in thermocycler at 42 C for 90 seconds
Place back on ice for 1 min
Place in incubator for 1 hour. 1126-1226
Plate cells on Spec plates and label them RDO 1214 3/15
Robert D O'Dowd 3/20
sbb1231
4 Colonies were picked by Zach.
Turns out none of the cell cultures were viable and the plates I created were totally unusable. There are 2 large red clusters where cells survived. The colonies that were picked were likely contamination from when I leaked. It is also possible that the transformation totally failed.
Experiment stopped. sbb1231 inconclusive.
sbb1214
4 colonies picked. All 4 cultures are viable.
Performing miniprep on all 4. [8]
Resulting solutions were labeled RO 1214 1, RO 1214 2, RO 1214 3, RO 1214 4
Running analytical digests on miniprepped plasmids
RO 1214 1 and 2
Digest with XbaI and NheI
Expect bands at 3161 and 456 if successful and 3161 and 1174 if unsuccessful
RO 1214 3 and 4
Digest with BamHI and EcoRI
Expect bands at 2472 and 1145 if successful and 2472 and 1863 if unsuccessful
Not enough time to incubate and run the gel.
Performing the gel on Thursday.
Removing 1231ROCT, 1231PCR3, 1231CITM, 1214Pca1, 1231TMPhoA from storage in order to make space.
Robert D O'Dowd 3/22
sbb1231
Not enough time to complete the experiment. I can redo the transformation and plating to at least figure out if my products were viable. If white colonies appear the Friday on a plate, I can say that the transformation at least happened.
Rerun ligation 1031-1101. pBca9525-Bca1834 digested with BamHI and EcoRI and ligated with RO 1231 Dig. Product is RO 1231 Lig2.
Competent cells thawed and prepared. Mixed with RO 1231 Lig2. Product is 1231 trans. Placed on ice for 10 min. 1113-1123
Incubate 1127-1127.
Cutting incubation 7 minutes early due to time.
Cells plated on Spec plates. Labeled RDO1231 3/22 No need to pick
sbb1214
Mapping miniprep
Thermocycler 30 min 1016-1046
Not sure if 1 and 2 will work. The XbaI enzyme solution is almost empty and less than .5 uL was added.
Run gel on lanes 1,2,3,4.
Lanes:
| 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 |
|---|---|---|---|---|---|---|---|---|---|
| none | RO 1214 1 | RO 1214 2 | RO 1214 3 | RO 1214 4 | 2-log | AJ digest | DC A+B | JW A10 | none |
All samples result in bands in the expected locations. Submitting 1214 2 and 1214 4 for sequencing.