SBB10Ntbk-XiaoY.Liu
Xiao Y. Liu 16:16, 8 March 2010 (EST)
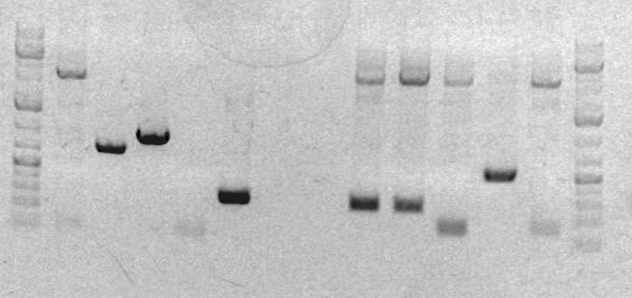
Xiao's gel on March 8th
Lane: 1. sbb03 2. PiggyBac 3. Ladder
PiggyBac
- after PCR 2, analytical gel (3ul PCR prodcut + 7ul loading buffer), look for 1813bp band
- Gel showed little SOEing products, so redo SOEing with 2k 45°C program.
SOEing
Set up the following reaction in a PCR tube:
24uL ddH2O 3.3uL 10x Expand Buffer "2" 3.3uL dNTPs (2mM in each) 1uL Oligo 1 = PiggyBac 01, 10uM 1uL Oligo 2 == PiggyBac 04, 10uM 0.5uL Expand polymerase "1" 0.5uL Template DNA from A+B+C stock solution
Sbb03
- after Amplification, run a analytical gel (3ul PCR prodcut + 7ul loading buffer),look for 550bp band
- failed, so give all products to prof.
Xiao Y. Liu 16:12, 3 March 2010 (EST)
PiggyBac
- proceed SOEing PCR after zymo gel purifications for all 3 parts(A, B, C)
- Elute the DNA products from all 3 gel purification reactions in 50uL of water, since i eluted each one with 8.5 ul, after combining everything, add 50-3*8.5 = 24.5 ul dd water.
- Set up your second round of PCR as a normal 33uL reaction using the eluted mixture of fragments as template
Set up the following reaction in a PCR tube:
24uL ddH2O 3.3uL 10x Expand Buffer "2" 3.3uL dNTPs (2mM in each) 1uL Oligo 1 = PiggyBac 01, 10uM 1uL Oligo 2 == PiggyBac 04, 10uM 0.5uL Expand polymerase "1" 0.5uL Template DNA from A+B+C stock solution
sbb03
- after re-assembly, proceed to second round of amplification.
Amplification
Now, you need to do an amplification of the correct full-length chunks. Clean up the assembly reaction with a zymo column; don't bother running it on a gel - it'll be a smeary mess and won't really help you.
Regular Zymo Cleanup
The following procedure removes the polymerase, dNTPs, buffer, and most of the oligonucleotides from a PCR reaction. It also will remove the buffer and restriction enzymes from a restriction digest reaction.
- Add 180 uL of Zymo ADB buffer (brown bottle) to a 33uL or 50uL reaction.
- Transfer into the Zymo column (small clear guys)
- spin through, discard waste.
- Add 200 uL of PE or Zymo Wash buffer (which is basically 70% ethanol)
- spin through, discard waste.
- Add 200 uL of PE or Zymo Wash buffer
- spin through, discard waste.
- spin for 90 seconds, full speed to dry.
- elute with water into a fresh Eppendorf tube, use the same volume of water as the volume of the original reaction (50ul in this case)
Save the purified product in case this step fails! For the amplification reaction, do a normal phusion program with 1 ul of the cleaned up assembly reaction as template, and using the outermost oligos for the chunk. That is:
Recipe
- 1 ul each outer oligo (10 uM)
- .5 ul phusion
- 10 ul 5x phusion buffer
- 5 ul 2mM dNTPs
- 32.5 ul H2O
Program
- 2 min initial denature at 94oC
- 30 sec denature at 94oC
- 30 sec anneal at 60oC [This should be high, as your outer oligos now have a huge overlap with the correct product]
- 30 sec extension at 68oC
- repeat 2-4 30 times total
Xiao Y. Liu 16:25, 1 March 2010 (EST)
PiggyBac
- Products from secomd PCR for piggyBac A, do a preparative gel with 6ul PCR products and 4ul loading buffer(blue, dye).
- prep gel showed success, so do a zymo gel purification for piggyBac A, eluted with 8.5ul water at the end. save stock and eluted DNA
sbb03
- Clean up the amplification reactions with a zymo column
Regular Zymo Cleanup
The following procedure removes the polymerase, dNTPs, buffer, and most of the oligonucleotides from a PCR reaction. It also will remove the buffer and restriction enzymes from a restriction digest reaction.
- Add 180 uL of Zymo ADB buffer (brown bottle) to a 33uL or 50uL reaction.
- Transfer into the Zymo column (small clear guys)
- spin through, discard waste.
- Add 200 uL of PE or Zymo Wash buffer (which is basically 70% ethanol)
- spin through, discard waste.
- Add 200 uL of PE or Zymo Wash buffer
- spin through, discard waste.
- spin for 90 seconds, full speed to dry.
- elute with water into a fresh Eppendorf tube, use the same volume of water as the volume of the original reaction (50ul)
- mistake: forgot to do a analytic gel, so use 6ul of eluted product with 4ul loading buffer to run a analytical gel.
- gel showed nothing, so do a re-assembly with 1 ul PCR product.
Xiao Y. Liu 16:21, 24 February 2010 (EST)
PiggyBac
Second PCR trial for PiggyBac A
- PCR for piggyBac A failed, so do a second PCR with oligos(ca1530 and piggyBac02) and with program 55.
make dilution(10uM) puggyBac01 from stock(100uM) with 9ul water and 1ul stock
set up PCR tube:
- lable tubes sbb04 A Xiao Liu 2/23/10
24uL ddH2O
3.3uL 10x Expand Buffer "2"
3.3uL dNTPs (2mM in each)
1uL Oligo 1, 10uM
1uL Oligo 2, 10uM
0.5uL Template DNA
0.5uL Expand polymerase "1"
- place them in 55 PCR program.
post preparative gel procedures for PiggyBac B and PiggyBac C
- after preparative gel, zymo purification.
Zymo Gel Purification
- All spins until the drying step are 15 second full speed spins.
- transfer into the Zymo column inside a collection tube (small clear guys)
- spin through, discard waste.
- Add 200 uL of PE buffer (which is basically 70% ethanol)
- spin through, discard waste.
- Add 200 uL of PE buffer
- spin through, discard waste.
- spin for 90 seconds, full speed to dry.
- elute with 8.5 uL of water into a fresh Eppendorf tube
Sbb03
Amplification reaction with a normal phusion program
- contiune sbb03 experiment. PCR with tos-1 and tos-20
- add following in order: (check order with Prof. )
1. 32.5 ul H2O 2. 10 ul 5x phusion buffer 3. 5 ul 2mM dNTPs 4. 1 ul each outer oligo (10 uM)=tos-1 and tos-20 5. 0.5 ul phusion
- run Phusion Program, anneal at 60°C, ~ 30 min.
Xiao Y. Liu 19:07, 22 February 2010 (EST)
PiggyBac
- sbb04 stops at melting gel with ADP buffer.
Sbb03
- sbb03 stops at the end of zymo clean up.
Image from preparative gel
1. ladder 2. sbb04A 3. sbb04B 4. sbb04C 5. RH 33A 6. RH33B 7. blank 8. blank 9. CD1 10. CD2 11. CD3 12. sbb19A 13. sbb19B 14. ladder
Xiao Y. Liu 16:40, 22 February 2010 (EST)
- After getting PCA products back for sbb03, do a zymo clean up, then do a PCR with tos-1 ans tos-20
- Products from 3 PCR for piggyBac, do a preparative gel with sbb04A, sbb04B, and sbb04 c with 6ul PCR products and 4ul loading buffer(blue, dye).
Xiao Y. Liu 03:37, 17 February 2010 (EST)
Modified protocol for Feb 17th, 2010
- set up PCA reaction for tos site. (no change in protocol)
- meanwhile, set up 3 PCR reactions for piggyBac.
- modified construction file for piggyBac
Construction file for piggyBac transposase (sbb04)
PCR with piggyBac01/CA1530 on Bca1519 3-2 (335bp, gp=A)
PCR CA1527/piggyBac03 on Bca1519 3-1 (721bp, gp=B)
PCR with piggyBac02/piggyBac04 on Bca1553 4-6 (925bp, gp=C)
-----------------------------------------------------------------
PCR with piggyBac01 and piggyBac04 on A+B+C (1813bp, EcoRI/BamHI)
Digest pBjk2741-Bca1144 (EcoRI/BamHI, 2170bp + 910bp,L)
Product is pBjk2741-sbb04 {<piggyBac!}
-----------------------------------------------------------------
piggyBac01 Forward PCR of part 1 of piggyBac
ccatagaattcatgAGATCTGGTTGCTCTCTGGACGACGAAC
CA1527 TGGACGAACAGAACGTTATCGAACAGCCGGGTTCTTCTCTGGCTTCT
PCA assembly of piggiebacfiveprime (Bca1519)
CA1530 ACGACGGGTAGATTTAGAGGTAGACCAGCAGTGTTTGTTTTTACCAC
PCA assembly of piggiebacfiveprime (Bca1519)
piggyBac02 Forward SOEing part of piggyBac
ACGGTATCAAAATCCTGATGatgtgcgactctggtaccaa
piggyBac03 Reverse SOEing part of piggyBac
ttggtaccagagtcgcacatCATCAGGATTTTGATACCGT
piggyBac04 Reverse PCR of part 2 of piggyBac
gctagGGATCCttaGAAGCAAGACTGGCACATG
- basic PCR method to amplify your DNA from a plasmid or genomic DNA sample using the Expand polymerase.
The oligo concentrations in my stocks should be 100uM. I use them at 10uM in this protocol. So, i first need to make an oligo dilution of:
9uL Water 1uL 100uM oligo
can throw away the remainder of the diluted oligo when i'm done, but hold onto the stock tube!
Set up the master mix in a PCR tube:
24uL ddH2O * 3 = 72 uL ddH2O
3.3uL 10x Expand Buffer "2" * 3 = 9.9uL 10x Expand Buffer "2"
3.3uL dNTPs (2mM in each) * 3 = 9.9 uL dNTPs
1.5uL Expand polymerase "1"
- lable tubes, divide the master mix into 3. and proceed following to each PCR reaction.
1uL Oligo 1, 10uM
1uL Oligo 2, 10uM
0.5uL Template DNA ( ask before this, Mike suggest to add this into master mix, but prof. said always add enzyme last, what should i do?)
- give GSI the tubes and run program.
Xiao Y. Liu 13:11, 16 February 2010 (EST)
Protocols in Order for sbb03
PCA
Assembly reaction
- add following materials in order. ~ 5 min
1.1. 38 uL ddH2O 1.2. 5 ul 10x expand buffer 1.3. 5 ul 2mM dNTPs 1.4. 1 ul oligo mixture (100uM total, mixture of oligos after combination of 100uM stocks) 1.5. 0.75 ul Expand polymerase
- run JCA/PCA1 program, (55°C for annealing) ~ 1.5 to 2 hours
- Zymo clean up to get rid of all the polymerase, dNTPS, extra oligos, and salt. ~5-10 min
The following procedure removes the polymerase, dNTPs, buffer, and most of the oligonucleotides from a PCR reaction. It also will remove the buffer and restriction enzymes from a restriction digest reaction. ( is there any changes in the buffers?)
1. Add 180 uL of Zymo ADB buffer (brown bottle) to a 33uL or 50uL reaction.
2. Transfer into the Zymo column (small clear guys)
3. spin through, discard waste.
4. Add 200 uL of PE or Zymo Wash buffer (which is basically 70% ethanol)
5. spin through, discard waste.
6. Add 200 uL of PE or Zymo Wash buffer
7. spin through, discard waste.
8. spin for 90 seconds, full speed to dry.
9. elute with water into a fresh Eppendorf tube, use the same volume of water as the volume of the original reaction
- save purified product, in case of failure.
Amplification reaction with a normal phusion program (~ 55 )
- add following in order: (check order with Prof. )
1. 32.5 ul H2O 2. 10 ul 5x phusion buffer 3. 5 ul 2mM dNTPs 4. 1 ul each outer oligo (10 uM) 5. 0.5 ul phusion
- run Phusion Program, anneal at 60°C, ~ 30 min.
- analytical gel, with 10 X dye, check length ~550bp,
- Zymo clean up to get rid of all the polymerase, dNTPS, extra oligos, and salt. ~3 min
The following procedure removes the polymerase, dNTPs, buffer, and most of the oligonucleotides from a PCR reaction. It also will remove the buffer and restriction enzymes from a restriction digest reaction. ( is there any changes in the buffers?)
1. Add 180 uL of Zymo ADB buffer (brown bottle) to a 33uL or 50uL reaction.
2. Transfer into the Zymo column (small clear guys)
3. spin through, discard waste.
4. Add 200 uL of PE or Zymo Wash buffer (which is basically 70% ethanol)
5. spin through, discard waste.
6. Add 200 uL of PE or Zymo Wash buffer
7. spin through, discard waste.
8. spin for 90 seconds, full speed to dry.
9. elute with water into a fresh Eppendorf tube, use the same volume of water as the volume of the original reaction
- digestion, mapping to make sure the length and sequence match. ~ 10 min
- if Fail, Re-assembly with the amplification product. repeat steps in Assembly reaction and Amplification reaction. IF succeed, move onto next step.
Cloning reaction ( ~ a few days )
- I have to get everything in vectors and clone the chunks together.
- Clean up the amplification reactions with a zymo column
- Digest the product and desired backbone with Eco/Bam ~ 5 min for step 1 and 2.
- Run everything on a gel, and cut out appropriate bands (~550bp) ~ 20 min
- Proceed with cloning as usual
- You should grow up, mini, and map 4 clones for gene synthesis, as we'll likely need to sequence more clones than we would for PCR-based part fabrication --- What does this mean?
Post PCA
- analytic gel
- Zymo clean up (>300bp)
- digestion
- gel purify, check if enough time. ( how long does gel purify take?)
Xiao Y. Liu 12:51, 16 February 2010 (EST)
- Planning for Experiment on Wed, Feb 17th, 2010
(sbb03) tos Site of N15 Flow Chart
- PCA with tos 1-tos 20, PCR with tos 1 and tos 20, 55°C.
- analytical gel to check length of products (550bp is the target sequence), cut out target band if find one.
- Zymo clean up
- digestion
- gel purify
- stop, check if enough time to proceed next step!!!
(sbb04) PiggyBac Transposase Flow Chart
- make master mix for 3 PCR reactions,
- PCR fragment A , B, and C
- analytic gel, check length
- Zymo clean up
- digestion, mapping?
- gel purify
- stop, check if enough time to proceed next step!!!
Xiao Y. Liu 18:30, 10 February 2010 (EST)
- check clotho, found that sbb04 DNA is not there, commit it.
- start tutorial 6
- correct mistakes in protein fusion construction files and ApE.
Xiao Y. Liu 18:26, 8 February 2010 (EST)
- modified construction file for both tos and piggyBac.
- upload sbb03 and sbb04 to clotho
- upload oligos to oligo log, and construction files.
Construction file for tos (modified)
1) sbb03: tos site of N15
Pool tos01 through tos20, assemble by PCA
PCR tos01/tos20 on PCA reaction (550bp,EcoR1/BamH1, gp)
Sub into pBca9145-jtk2625 (EcoRI/BamHI, 2057+929bp, L)
Product is pBca9145-sbb03 {<tos>}
------------
tos01 PCA assembly of tos CCATAGAATTCATGAGATCTTCTAAGCGCAACGGTATTACTTACGT
tos02 PCA assembly of tos ATTTAAAATCATTAAGTTAGGTTTTAAATATACCAACGTAAGTAATACCGTTGC
tos03 PCA assembly of tos TTAAAACCTAACTTAATGATTTTAAATGATAATAAATCATACCAATTGC
tos04 PCA assembly of tos CAGCATGTTCGCTTAACTTTTGATAGCAATTGGTATGATTTATTATC
tos05 PCA assembly of tos AAAGTTAAGCGAACATGCTGATTTTCACGCTGTTTATACACTTTGAG
tos06 PCA assembly of tos ATATAGAGACGGAAGAGATAGAGATGCCTCAAAGTGTATAAACAGCG
tos07 PCA assembly of tos TATCTCTTCCGTCTCTATATTGAAACACAATCAAAGAACATCAATCC
tos08 PCA assembly of tos TTCTTAGATAGTGGGGGATGTCACATGGATTGATGTTCTTTGATTG
tos09 PCA assembly of tos ATCCCCCACTATCTAAGAACACCATAACAGAACACAACATAGGAATG
tos10 PCA assembly of tos TGTTCCGAATTATTGATACATTAATGTTGCATTCCTATGTTGTGTTCTG
tos11 PCA assembly of tos AATGTATCAATAATTCGGAACATATGCACTATATCATATCTCAATTACGG
tos12 PCA assembly of tos ATGGGCAATTGTGTGCTGATATGTTCCGTAATTGAGATATGATATAG
tos13 PCA assembly of tos TCAGCACACAATTGCCCATTATACGCGCGTATAATGGACTATTGTGTG
tos14 PCA assembly of tos GTTCTGCGCTTATGTTCTCCTTATCAGCACACAATAGTCCATTATAC
tos15 PCA assembly of tos GAGAACATAAGCGCAGAACAATATGTATCTATTCCGGTGTTGTGTTC
tos16 PCA assembly of tos TATAAGAGAACATAATAGCAGAATAACAAAGGAACACAACACCGGAATAG
tos17 PCA assembly of tos TCTGCTATTATGTTCTCTTATAGTGTGACGAAAGCAGCATAATTAATC
tos18 PCA assembly of tos ATATCGTAACACAATCAAAGAACAAGTGACGATTAATTATGCTGCTTTCG
tos19 PCA assembly of tos TTCTTTGATTGTGTTACGATATCCAGAGACTTAGAAACGGGGGAAGG
tos20 PCA assembly of tos CATAGGGATCCTTCCCCCGTTTCTAAGT
Construction file for piggyBac transposase (modified)
Construction file for piggyBac transposase (sbb04)
PCR with piggyBac01/CA1530 on Bca1519 3-2 (335bp, gp=A)
PCR CA1527/piggyBac03 on Bca1519 3-1 (721bp, gp=B)
PCR with piggyBac02/piggyBac04 on Bca1553 4-6 (925bp, gp=C)
-----------------------------------------------------------------
PCR with piggyBac01 and piggyBac04 on A+B+C (1813bp, EcoRI/BamHI)
Digest pBjk2741-Bca1144 (EcoRI/BamHI, 2170bp + 910bp,L)
Product is pBjk2741-sbb04 {<piggyBac!}
-----------------------------------------------------------------
piggyBac01 Forward PCR of part 1 of piggyBac
ccatagaattcatgAGATCTGGTTGCTCTCTGGACGACGAAC
CA1527 TGGACGAACAGAACGTTATCGAACAGCCGGGTTCTTCTCTGGCTTCT
PCA assembly of piggiebacfiveprime (Bca1519)
CA1530 ACGACGGGTAGATTTAGAGGTAGACCAGCAGTGTTTGTTTTTACCAC
PCA assembly of piggiebacfiveprime (Bca1519)
piggyBac02 Forward SOEing part of piggyBac
ACGGTATCAAAATCCTGATGatgtgcgactctggtaccaa
piggyBac03 Reverse SOEing part of piggyBac
ttggtaccagagtcgcacatCATCAGGATTTTGATACCGT
piggyBac04 Reverse PCR of part 2 of piggyBac
gctagGGATCCttaGAAGCAAGACTGGCACATG
Xiao Y. Liu 15:50, 8 February 2010 (EST)
PiggyBac
- propose a construction file for sbb04, piggyBac transposase,
- question about whether to leave a start codon in the beginning of sequence. since the falimy flavor is {<part!}, so i leave it out.
2) construction file for piggyBac transposase (sbb04)
PCR with piggyBac01 and piggyBac03 on pBca9523-Bca1519 (928bp, gp, A)
PCR with piggyBac02 and piggyBac04 on pBca9523-Bca1553 (925bp, gp, B)
-----------------------------------------------------------------
PCR with piggyBac01 and piggyBac04 on A+B (1813bp, EcoR1/BamH1)
Digest pBjk2741-Bca1144 (EcoR1/BamH1, 2170bp + 910bp,L)
Product is pBjk2741-sbb04 {<piggyBac!}
-----------------------------------------------------------------
piggyBac01 Forward PCR of part 1 of piggyBac ccatagaattcatgAGATCTGGTTGCTCTCTGGACGACGAAC
piggyBac02 Forward SOEing part of piggyBac ACGGTATCAAAATCCTGATGatgtgcgactctggtaccaa
piggyBac03 Reverse SOEing part of piggyBac ttggtaccagagtcgcacatCATCAGGATTTTGATACCGT
piggyBac04 Reverse PCR of part 2 of piggyBac gctagGGATCCttaGAAGCAAGACTGGCACATG
- description of PCR product: ccata + EcoR1 + atg + bgl2 + part 1 (last 20bp + first 20bp as SOEing Part) and part 2 of piggyBac + stop codon + BamH1 + ctagc.
Xiao Y. Liu 20:00, 6 February 2010 (EST)
Sbb03 tos
oligo design for {<tos>} using geneDesign
- 1 biolding block, length of building block is 550 bp,
- average Tm of oligo overlaps in building block is 46°,
- number of oligos in building block: 20,
- average oligo length: 47bp, longest oligo: 54bp, shorest oligo: 28bp.
- PCA product is 5 random + EcoR1 + atg + bgl2 + target sequence + BamH1 + 5 random
Construction file for tos (check for errors)
1) sbb03: tos site of N15 Pool tos01 through tos20, assemble by PCA PCR tos01/tos20 on PCA reaction (550bp,EcoR1/BamH1, gp) Sub into pBca9145-jtk2625 (EcoRI/BamHI, 2057+929bp, L) Product is pBca9145-sbb03 ------------ tos01 PCA assembly of tos CCATAGAATTCATGAGATCTTCTAAGCGCAACGGTATTACTTACGT tos02 PCA assembly of tos ATTTAAAATCATTAAGTTAGGTTTTAAATATACCAACGTAAGTAATACCGTTGC tos03 PCA assembly of tos TTAAAACCTAACTTAATGATTTTAAATGATAATAAATCATACCAATTGC tos04 PCA assembly of tos CAGCATGTTCGCTTAACTTTTGATAGCAATTGGTATGATTTATTATC tos05 PCA assembly of tos AAAGTTAAGCGAACATGCTGATTTTCACGCTGTTTATACACTTTGAG tos06 PCA assembly of tos ATATAGAGACGGAAGAGATAGAGATGCCTCAAAGTGTATAAACAGCG tos07 PCA assembly of tos TATCTCTTCCGTCTCTATATTGAAACACAATCAAAGAACATCAATCC tos08 PCA assembly of tos TTCTTAGATAGTGGGGGATGTCACATGGATTGATGTTCTTTGATTG tos09 PCA assembly of tos ATCCCCCACTATCTAAGAACACCATAACAGAACACAACATAGGAATG tos10 PCA assembly of tos TGTTCCGAATTATTGATACATTAATGTTGCATTCCTATGTTGTGTTCTG tos11 PCA assembly of tos AATGTATCAATAATTCGGAACATATGCACTATATCATATCTCAATTACGG tos12 PCA assembly of tos ATGGGCAATTGTGTGCTGATATGTTCCGTAATTGAGATATGATATAG tos13 PCA assembly of tos TCAGCACACAATTGCCCATTATACGCGCGTATAATGGACTATTGTGTG tos14 PCA assembly of tos GTTCTGCGCTTATGTTCTCCTTATCAGCACACAATAGTCCATTATAC tos15 PCA assembly of tos GAGAACATAAGCGCAGAACAATATGTATCTATTCCGGTGTTGTGTTC tos16 PCA assembly of tos TATAAGAGAACATAATAGCAGAATAACAAAGGAACACAACACCGGAATAG tos17 PCA assembly of tos TCTGCTATTATGTTCTCTTATAGTGTGACGAAAGCAGCATAATTAATC tos18 PCA assembly of tos ATATCGTAACACAATCAAAGAACAAGTGACGATTAATTATGCTGCTTTCG tos19 PCA assembly of tos TTCTTTGATTGTGTTACGATATCCAGAGACTTAGAAACGGGGGAAGG tos20 PCA assembly of tos CATAGGGATCCTTCCCCCGTTTCTAAGT