SBB10Ntbk-AndrewSaarni: Difference between revisions
(→sbb12) |
|||
| Line 242: | Line 242: | ||
After thermocycling, a mixture of 6uL of product and 4uL of loading buffer was run on a preparative gel.<br> | After thermocycling, a mixture of 6uL of product and 4uL of loading buffer was run on a preparative gel.<br> | ||
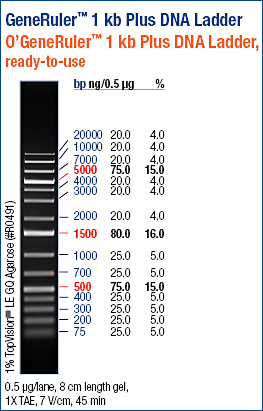
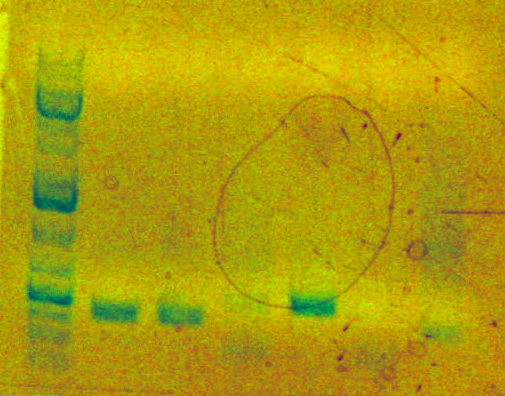
[[Image:Generuler1kbplus.jpg]][[Image:AS22610.png]] | [[Image:Generuler1kbplus.jpg]][[Image:AS22610.png]] | ||
Lane 7 = AS04 (259bp) | Lane 7 = AS04 (259bp)<br> | ||
The appropriate band was then cut out of the gel and resuspended at 55oC in 600uL of ADB buffer for weekend storage.<br> | The appropriate band was then cut out of the gel and resuspended at 55oC in 600uL of ADB buffer for weekend storage.<br> | ||
===sbb14=== | ===sbb14=== | ||
Revision as of 16:34, 26 February 2010
Design
Construction files for two parts (Sleeping Beauty 3'TR and PhiC31 Integrase with RBS)
sbb12
The sbb12 part is small enough to be entirely synthesized using PCA. The GeneDesign program [1]
was used to generate appropriate oligos for the synthesis.
1) sbb12: Sleeping Beauty 3'TR ---------------------------------------------------------- Pool AS001-F through AS008-R, assemble by PCA PCR AS001-F/AS008-R on PCA reaction (259 bp, EcoRI/BamHI) Substitute in pBjk2741-Bca1144 (EcoRI/BamHI, 910+2170, L) Product is pBjk2741-sbb12 ---------------------------------------------------------- AS001-F PCA Assembly of sbb12 CCATAGAATTCATGAGATCTTTGAGTGTATGTTAACTTCTGACCCACTGG AS002-R PCA Assembly of sbb12 ATTTCAGCTTTTATTTCTTTCATCACATTCCCAGTGGGTCAGAAGTTAAC AS003-F PCA Assembly of sbb12 AAAGAAATAAAAGCTGAAATGAATCATTCTCTCTACTATTATTCTGATAT AS004-R PCA Assembly of sbb12 AGGATCACCACTTTATTTTAAGAATGTGAAATATCAGAATAATAGTAGAG AS005-F PCA Assembly of sbb12 TAAAATAAAGTGGTGATCCTAACTGACCTTAAGACAGGGAATCTTTACTC AS006-R PCA Assembly of sbb12 CACTTTTTCACAATTCCTGACATTTAATCCGAGTAAAGATTCCCTGTCTT AS007-F PCA Assembly of sbb12 TCAGGAATTGTGAAAAAGTGAGTTTAATGTATTTGGCTAAGGTGTATGTA AS008-R PCA Assembly of sbb12 CTGATGGATCCCAGTTGAAGTCGGAAGTTTACATACACCTTAGCCAAAT
sbb14
For construction, sbb14 will be created as three sections which will then be connected by SOEing PCR.
Each section is constructed by amplifying a template DNA from a plasmid using standard PCR amplification.
The resulting sections have complementary overlapping sequences (either at the beginning, end, or both),
which allow the sections to be attached to each other by SOEing PCR to produce the final part.
Additionally, a strong RBS was generated using [2] and added to the first
forward oligo to be incorporated into the first section.
sbb14: PhiC31 Integrase
----------------------------------------------------------
PCR AS009-F/AS010-R on Bca1623 6-9 (976bp, gp=A)
PCR AS011-F/CA1674 on Bca1559 7-7 (514bp, gp=B)
PCR CA1675/AS012-R on Bca1559 7-1 (444bp, gp=C)
----------------------------------------------------------
PCR AS009-F/AS012-R on A+B+C (1868 bp, EcoRI/BamHI)
Sub in pBjk2741-Bca1144 (EcoRI/BamHI, 910+2170, L)
Product is pBjk2741-sbb14 {rbs.phiC31>}
----------------------------------------------------------
AS009-F Forward PCR of Part 1 of PhiC31 with RBS ccataGAATTCatgAGATCTCAGATCATATATAAGGAGGTACATatgGACACGTACGCGGGTGCTTACG
AS010-R SOEing of Part 1 of PhiC31 cggtaaccctcaatcttcgtGGTCGGCGTGCCGTCCGGCTTCTT
AS011-F SOEing of Part 2 of PhiC31 AAGAAGCCGGACGGCACGCCGACCacgaagattgagggttaccg
CA1674 PCA assembly of phiCthreeprime (Bca1659) GGGCGTCGGCGCGCTCCGCAACAAGGTTCGCCCGTTCGCCGCTCTTCTCAG
CA1675 PCA assembly of phiCthreeprime (Bca1659) TGCGGAGCGCGCCGACGCCCTGAACGCCCTTGAAGAGCTGTACGAAGACCG
AS012-R Reverse PCR of Part 2 of PhiC31 atcagGGATCCCGCCGCTACGTCTTCCGT
Part: GATCTCAGATCATATATAAGGAGGTACATatgGACACGTACGCGGGTGCTTACGACCGTCAGTCGCGCGAGCGCGAAAATTCGAGCGCAGCAAGCCCAGCGACACAGCGTAGCGCCAACGAAGACAAGGCGGCCGACCTCCAGCGCGAAGTCGAGCGCGACGGGGGCCGGTTCAGGTTCGTCGGGCATTTCAGCGAAGCGCCGGGCACGTCGGCGTTCGGGACGGCGGAGCGCCCGGAGTTCGAACGCATCCTGAACGAATGCCGCGCCGGGCGGCTCAACATGATCATTGTCTATGACGTGTCGCGCTTCTCGCGCCTCAAGGTCATGGACGCGATTCCGATTGTCTCGGAATTGCTCGCCCTGGGCGTGACGATTGTTTCCACTCAGGAAGGCGTCTTCCGGCAGGGAAACGTCATGGACCTGATTCACCTGATTATGCGGCTCGACGCGTCGCACAAAGAATCTTCGCTCAAGTCGGCGAAGATTCTCGACACGAAGAACCTCCAGCGCGAATTGGGCGGGTACGTCGGCGGGAAGGCGCCTTACGGCTTCGAGCTTGTTTCGGAGACGAAGGAGATCACGCGCAACGGCCGAATGGTCAATGTCGTCATCAACAAGCTTGCGCACTCGACCACTCCCCTTACCGGACCCTTCGAGTTCGAGCCCGACGTAATCCGGTGGTGGTGGCGTGAGATCAAGACGCACAAACACCTTCCCTTCAAGCCGGGCAGTCAAGCCGCCATTCACCCGGGCAGCATCACGGGGCTTTGTAAGCGCATGGACGCTGACGCCGTGCCGACCCGGGGCGAGACGATTGGGAAGAAGACCGCTTCAAGCGCCTGGGACCCGGCAACCGTTATGCGAATCCTTCGGGACCCGCGTATTGCGGGCTTCGCCGCTGAGGTGATCTACAAGAAGAAGCCGGACGGCACGCCGACCacgaagattgagggttaccgCATTCAGCGCGACCCGATCACGCTCCGGCCGGTCGAGCTTGATTGCGGACCGATCATCGAGCCCGCTGAGTGGTATGAGCTCCAGGCGTGGTTGGACGGCAGGGGGCGCGGCAAGGGGCTTTCCCGGGGGCAAGCCATTCTGTCCGCCATGGACAAGCTGTACTGCGAGTGTGGCGCCGTCATGACTTCGAAGCGCGGGGAAGAATCGATCAAGGACTCTTACCGCTGCCGTCGCCGGAAGGTGGTCGACCCGTCCGCACCTGGGCAGCACGAAGGCACGTGCAACGTCAGCATGGCGGCACTCGACAAGTTCGTTGCGGAACGCATCTTCAACAAGATCAGGCACGCCGAAGGCGACGAAGAGACGTTGGCGCTTCTGTGGGAAGCCGCCCGACGCTTCGGCAAGCTCACTGAGGCGCCTGAGAAGAGCGGCGAACGGGCGAACCTTGTTGCGGAGCGCGCCGACGCCCTGAACGCCCTTGAAGAGCTGTACGAAGACCGCGCGGCAGGCGCGTACGACGGACCCGTTGGCAGGAAGCACTTCCGGAAGCAACAGGCAGCGCTGACGCTCCGGCAGCAAGGGGCGGAAGAGCGGCTTGCCGAACTTGAAGCCGCCGAAGCCCCGAAGCTTCCCCTTGACCAATGGTTCCCCGAAGACGCCGACGCTGACCCGACCGGCCCTAAGTCGTGGTGGGGGCGCGCGTCAGTAGACGACAAGCGCGTGTTCGTCGGGCTCTTCGTAGACAAGATCGTTGTCACGAAGTCGACTACGGGCAGGGGGCAGGGAACGCCCATCGAGAAGCGCGCTTCGATCACGTGGGCGAAGCCGCCGACCGACGACGACGAAGACGACGCCCAGGACGGCACGGAAGACGTAGCGGCGG
Assembly
2.17.2010
Overall Progress
Performed initial step of PCA assembly of sbb12 and initial step of SOEing PCR for sbb14.
sbb12 progress: Design --> Assembly --> Amplification --> Reassembly --> Cloning
sbb14 progress: Initial PCR Reactions --> Zymo Clean-up --> Secondary PCR Reactions --> Cloning
sbb12
Experimental
Oligo Mixture was provided at 100uM
Initial assembly of the PCA oligo mixture was performed according to the following protocol:
1. 38 uL ddH2O 2. 5 ul 10x expand buffer 3. 5 ul 2mM dNTPs 4. 1 ul oligo mixture (100uM total, mixture of oligos after combination of 100uM stocks) 5. 0.75 ul Expand polymerase
The resulting mixture was then thermo-cycled according to the following program:
1. 2 min initial denature at 94oC 2. 30 sec denature at 94oC 3. 30 sec anneal at 55oC [This should be the overlap temp of your oligos - vary as needed] 4. 30 sec extension at 72oC 5. repeat 2-4 30 times total
Note: thermo-cycling performed by GSI's
sbb14
Experimental
All oligo samples were first diluted to 100uM by adding sufficient water (ddH2O)
Then, 10uM samples of each oligo were prepared by combining 1uL of 100uM Oligo and 9uL of ddH2O
Three PCR tubes were labeled 1,2 and 3.
The following PCR protocol was used:
24uL ddH2O 3.3uL 10x Expand Buffer "2" 3.3uL dNTPs (2mM in each) 1uL Oligo 1, 10uM 1uL Oligo 2, 10uM 0.5uL Expand polymerase "1" 0.5uL Template DNA
A forward and reverse oligo were added to each tube according to the PCR protocol.
Tube 1: AS009-F and AS010-R
Tube 2: AS011-F and CA1674
Tube 3: CA1675 and AS012-R
Tubes 1,2 and 3 were then thermo-cycled according to the 2K55 program because each PCR product
is less than 2000bp.
2.22.2010
Overall Progress
Ran preparative gel for sbb14 SOEing reactions and performed a small-frag Zymo Cleanup followed by Fuzion amplification for sbb12 PCA
sbb12 progress: Design --> Assembly --> Amplification --> Reassembly --> Cloning
sbb14 progress: Initial PCR Reactions --> Preparative Gel --> Zymo Clean-up --> Secondary PCR Reactions
sbb12
Experimental
Initial PCA Assembly reaction was purified by small fragment Zymo cleanup according to the following protocol:
Small-Frag Zymo Cleanup The following procedure removes the polymerase, dNTPs, buffer, and most of the oligonucleotides from a PCR reaction for fragments smaller than 300bp. It also will remove the buffer and restriction enzymes from a restriction digest reaction. 1. Add 100 uL of Zymo ADB buffer (brown bottle) to the reaction. 2. Transfer into the Zymo column (small clear guys) 3. Add 500uL of Ethanol and pipette up and down to mix 4. spin through, discard waste. 5. Add 200 uL of PE or Zymo Wash buffer (which is basically 70% ethanol) 6. spin through, discard waste. 7. Add 200 uL of PE or Zymo Wash buffer 8. spin through, discard waste. 9. spin for 90 seconds, full speed to dry. 10. elute with water into a fresh Eppendorf tube
The purified product was used as the template for the following PCR amplification protocol:
Amplification Now, you need to do an amplification of the correct full-length chunks. Clean up the assembly reaction with a zymo column; don't bother running it on a gel - it'll be a smeary mess and won't really help you. Save the purified product in case this step fails! For the amplification reaction, do a normal phusion program with 1 uL of the cleaned up assembly reaction as template, and using the outermost oligos for the chunk. That is: Recipe 1. 1 uL each outer oligo (10 uM) (Provided as F/R Oligos) 2. .5 uL phusion 3. 10 uL 5x phusion buffer 4. 5 uL 2mM dNTPs 5. 32.5 uL H2O 6. 1 uL Template Program 1. 2 min initial denature at 94oC 2. 30 sec denature at 94oC 3. 30 sec anneal at 60oC [This should be high, as your outer oligos now have a huge overlap with the correct product] 4. 30 sec extension at 68oC 5. repeat 2-4 30 times total
Tube labeled AS Fuzion 04
sbb14
Experimental
6uL of each PCR reaction and 4uL of loading buffer were added to 3 Eppendorf tubes, labeled AS01, AS02 and AS03.
Resulting mixtures were then run on preparative gels.

Lane 3 = AS01 (976bp)
Lane 4 = AS02 (514bp
Lane 5 = AS03 (444bp)
Bands are at correct sizes; the PCR reaction was successful.
The three bands were cut out, placed in a 1.5mL microcentrifuge tube, and resuspended in 600uL of ADB buffer.
PCR product mixture was stored until 2.24.2010.
2.24.2010
Overall Progress
PCA Amplification reaction for sbb12 was successful and the product was purified using Small-Frag Zymo Cleanup.
Performed Zymo Gel Purification and second PCR reaction for sbb14
sbb12 progress: Design --> Assembly --> Amplification --> Zymo Clean-up --> Restriction Digest --> Preparative Gel --> Cloning
sbb14 progress: Initial PCR Reactions --> Preparative Gel --> Zymo Gel Purification --> Secondary PCR Reactions--> Analytical Gel --> Restriction Digest --> Cloning
sbb12
Analytical gel was run for PCA amplification product.

Lane 4 = AS04 (259bp)
Smearing is expected with PCA, but a band nevertheless appears at the correct size.
Amplification was deemed a success and a Small-Frag Zymo Cleanup was performed on the amplification product.
Restriction digest with EcoR1/BamH1 will be performed Friday, 2.26.2010.
sbb14
PCR Product mixture was purified using Small-Frag Zymo Cleanup.
Purified fragments were eluted in 50uL of ddH2O.
Tube was labeled AS123.
Second round of PCR was then set up using the purified mixture of fragments as template DNA.
AS009-F and AS012-R were used as the oligos for the PCR.
The following PCR protocol was used:
24uL ddH2O 3.3uL 10x Expand Buffer "2" 3.3uL dNTPs (2mM in each) 1uL Oligo 1, 10uM (AS009-F) 1uL Oligo 2, 10uM (AS012-R) 0.5uL Expand polymerase "1" 0.5uL Template DNA (AS123)
2.26.2010
Overall Progress
Digested sbb12 with EcoR1/BamH1 and ran the product on a preparative gel.
Ran product of second round of PCR for sbb14 on an analytical gel.
sbb12 progress: Design --> Assembly --> Amplification --> Zymo Clean-up --> Restriction Digest --> Preparative Gel --> Cloning
sbb14 progress: Initial PCR Reactions --> Preparative Gel --> Zymo Gel Purification --> Secondary PCR Reactions --> Analytical Gel --> Restriction Digest --> Cloning
sbb12
Performed EcoR1/BamH1 restriction digest on cleaned up PCA amplification product according to the following protocol:
8uL of eluted PCR product 1uL of NEB Buffer 2 0.5uL EcoRI 0.5uL BamHI
Reaction mixture was then placed in a thermocycler at 37C for 1 hour.
After thermocycling, a mixture of 6uL of product and 4uL of loading buffer was run on a preparative gel.

 Lane 7 = AS04 (259bp)
Lane 7 = AS04 (259bp)
The appropriate band was then cut out of the gel and resuspended at 55oC in 600uL of ADB buffer for weekend storage.
sbb14
Product of the second round of PCR was run on an analytical gel.

 Lane 8 = AS123 (1868bp)
Lane 8 = AS123 (1868bp)
No band was observed in Lane 8. SOEing failed.
Restriction digest will be performed Monday, 3.1.2010.