IGEM:IMPERIAL/2006/project/popsblocker/Implementation: Difference between revisions
From OpenWetWare
Jump to navigationJump to search
No edit summary |
|||
| Line 12: | Line 12: | ||
[[Image:Cre.PNG]] | [[Image:Cre.PNG]] | ||
==Ligation strategy== | |||
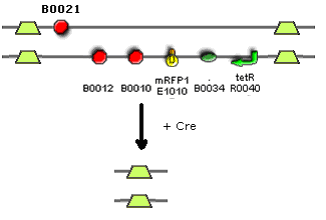
*Apart from ligating into the necessary plasmid at the end, there are no ligation steps involved in making this device. PCR and PCR fusion are being are used because of the unusual design of the device, in so far as both strands have coding regions. The link below explains the method by which we will use PCR to create the device. | |||
[[Image:Cre3.PNG]] | [[Image:Cre3.PNG]] | ||
*[http://openwetware.org/images/5/5a/Cre_Pops_Blocker.ppt Proposed Implementation of Cre-LoxP system (ppt)] | |||
==Considerations== | |||
'''Stop Codons''' | |||
A stop codon has been included between Lox66 and B0021 to prevent the ''E.Coli'' cells from translating past the first Lox site. This reduces the load that this part will place on the E.coli cell so increases its stability ''in vivo''. TGA is by far the most commonly used stop codon in ''E.Coli''. | |||
'''Frameshift''' | |||
The Lox sites are 34bp long therefore if this part is placed after an RBS it will induce a frameshift in the protein produced, therefore creating a mangled protein. Two random bases were added to the start of the Lox site to prevent a frameshift. | |||
==Primers== | ==Primers== | ||
[http://openwetware.org/images/5/ | [http://openwetware.org/images/5/58/Redoing_Cre2a.doc Design of primers and method of construction] | ||
[http://openwetware.org/images/b/ba/A112-705-Cre_Version_2_1.pdf Commercially available Cre plasmid] | [http://openwetware.org/images/b/ba/A112-705-Cre_Version_2_1.pdf Commercially available Cre plasmid] | ||
==WetLab work entries== | ==WetLab work entries== | ||
Revision as of 13:50, 29 October 2006
Ligation strategy
- Apart from ligating into the necessary plasmid at the end, there are no ligation steps involved in making this device. PCR and PCR fusion are being are used because of the unusual design of the device, in so far as both strands have coding regions. The link below explains the method by which we will use PCR to create the device.
Considerations
Stop Codons
A stop codon has been included between Lox66 and B0021 to prevent the E.Coli cells from translating past the first Lox site. This reduces the load that this part will place on the E.coli cell so increases its stability in vivo. TGA is by far the most commonly used stop codon in E.Coli.
Frameshift
The Lox sites are 34bp long therefore if this part is placed after an RBS it will induce a frameshift in the protein produced, therefore creating a mangled protein. Two random bases were added to the start of the Lox site to prevent a frameshift.
Primers
Design of primers and method of construction
Commercially available Cre plasmid
WetLab work entries
Jonny Wells <br\> Kirsten Jensen
(08/06 - 10/06)