CH391L/S13/In vitro Selection of FNAs: Difference between revisions
No edit summary |
No edit summary |
||
| Line 7: | Line 7: | ||
==Functional Nucleic Acids== | ==Functional Nucleic Acids== | ||
<cite>Cech1982</cite> | |||
==In vitro Selection of Functional Nucleic Acids== | ==In vitro Selection of Functional Nucleic Acids== | ||
| Line 31: | Line 31: | ||
Chemical synthesis is currently limited to oligonucleotides of about 200 nt in length. | Chemical synthesis is currently limited to oligonucleotides of about 200 nt in length. | ||
#Cech1982 pmid=6297745 | |||
Revision as of 21:46, 10 February 2013
Introduction
Functional nucleic acids (FNAs) are RNA, DNA, or XNA(nucleic acid analogues) that perform an activity such as binding or catalyzing a reaction. FNAs are grouped into three main categories Aptamers, Ribozymes, and Deoxyribozymes that are subdivided into either natural or artificial depending on their origin; the exception being Deoxyribozymes as they have yet to be discovered in a living organism.
Functional Nucleic Acids
[1]
In vitro Selection of Functional Nucleic Acids
Ribozymes
Deoxyribozymes
Extra
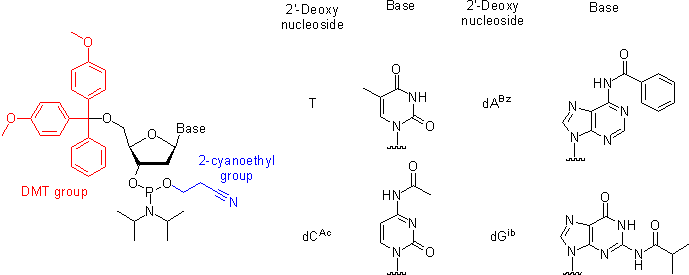
Oligonucleotides are chemically synthesized from DNA phosphoramidite monomers. Briefly, activated phosphoramidite monomers are added in the 3' to 5' direction using a cyclical activation and blocking chemistry to obtain a DNA polymer linked by phosphodiester bonds.
Chemical synthesis is currently limited to oligonucleotides of about 200 nt in length.
- Cech1982 pmid=6297745