CH391L/S13/DnaAssembly
As described before, synthetic biology captures a diverse, multi-disciplinary field. No matter which definition(s) becomes accepted, the ability to make and manipulate DNA is a vital component to practicing synthetic biology.
A large number of parts have been made by the synthetic biology community. Many can be found as part of the Registry of Standard Biological Parts. These modular genetic components are designed to be easy to acquire and assemble to facilitate the building of more complex biological devices. To learn more about the Registry and the biological parts known as BioBricks™, see the entry for the iGEM Registry.
DNA Synthesis
Oligonucleotide Synthesis
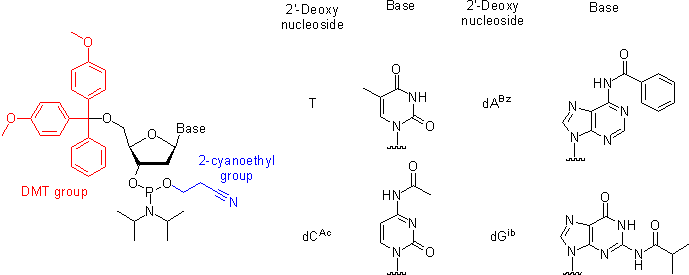
Oligonucleotides, short single-stranded DNA pieces, are chemically synthesized from DNA phosphoramidite monomers. Briefly, activated phosphoramidite monomers are added in the 3' to 5' direction using a cycle of activation and blocking chemistry to obtain a DNA polymer linked by phosphodiester bonds.
Chemical synthesis is currently limited to oligonucleotides of about 200 nt in length. It is used often by synthetic biologists to make primers for PCR or gibson assembly (described later).
Gene Synthesis
Gene synthesis, or artificial gene synthesis, refers to the process of creating a nucleic acid template for a gene in vitro, without the requirement of a preexisting DNA template. It predates the invention of restriction enzymes and molecular cloning techniques by several years. The first gene to be completely synthesized in vitro was a 77-nt alanine transfer RNA by the laboratory of Har Gobind Khorana in 1972 [1]. This was the result of nearly five years of work and resulted in a DNA template without promoter or transcriptional control sequences. The first peptide- and protein-producing synthetic genes were created in 1977 and 1979, respectively [2, 3]. Chemical synthesis of DNA has grown from an expensive and time-consuming process into a viable commercial industry capable of versatile and high-throughput manufacturing. DNA can now be made to order for species-specific gene optimization, creation of genes from rare or dangerous sources, and combinatorial assembly of any DNA sequence that can be chemically synthesized, including non-natural bases.

Despite nearly four decades of progress in gene synthesis technologies, most DNA sequences used in modern molecular biology are assembled in part or in whole from naturally occurring templates. However this limits the scope and applications to previously existing genes and the results of large-scale genomic surveys of novel genes from nature. Modern gene synthesis relies heavily on advancements in chemical DNA oligonucleotide synthesis, with the primary challenges being scale, cost, fidelity and the eventual assembly of complete gene products.
An extensive, but not comprehensive, directory of commercial gene synthesis providers can be found at Genespace.
Economics
Gene synthesis is one of the key enabling technologies of synthetic biology. The increase in efficiency and the decrease in cost is occurring at a staggering rate. Below is a graph comparing DNA synthesis to Moore's Law. In addition, it includes another enabling technology, DNA sequencing. The bottom right graph shows that genome synthesis is technically feasible [4]; however, it is not yet a commercial service.


Molecular Cloning
It is impractical for most synthetic biologists to synthesize more than several kilobases of completely synthetic DNA. It is often desirable to build bigger pieces at lower costs and faster speeds than de novo synthesis is currently able to accomplish.
Over the years, many different strategies have been developed to assemble DNA in flexible ways that suit different purposes. These strategies typically employ purification of enzymes that are known to modify DNA in specific ways and these methods of action can be exploited for designing and building specific sequences of DNA. For example, restriction enzymes are used to cut DNA in a specific manner upon recognition of a specific nucleotide sequence. Polymerases and endonucleases add or remove nucleotides to make double stranded DNA from a single stranded template or to create single stranded DNA from double stranded DNA. Many of the modern techniques take advantage of recombination machinery that move DNA from one location to another location. In other cases, the endogenous enzymes in a host are utilized to manipulate DNA without the need for prior purification. For example, in some methods endogenous DNA ligase is used to repair single stranded breaks (known as "nicks") to complete the formation of fully circular DNA.
Ultimately, these methods generally require transformation into a host where endogenous enzymes are used to complete the genetic manipulation and replicate (clone) the genetic material. This allows for the expression of the desired proteins to test the ability of the engineered system or for the purification of the genetic material itself such that it can be used for further manipulation, study, or storage.
While there are many specific protocols for the numerous methods of cloning, most share reasonable overlap in their underlying mechanisms of action. Broadly speaking, methods may rely primarily on restriction enzymes, polymerase chain reaction (PCR), or on homologous recombination [5, 6].
Restriction Enzyme
Restriction enzymes recognize a specific nucleotide sequence and then cut the DNA in such a way that results in a double stranded break. If the enzyme cuts within or near the recognition site, it is classified as a Type II restriction enzyme. When a restriction enzyme cuts a piece of DNA, the resulting ends can be straight resulting in a blunt end or with a jagged cut resulting in a sticky end.
For example:
EcoRI digestion produces "sticky" ends,

whereas AluI restriction enzyme cleavage produces "blunt" ends:

The Registry of Biological Parts has developed standards for the type and position of restriction enzyme sites to be used when building DNA using restriction enzymes in order to ensure compatibility in the connecting of biological parts. Two of the more popular standards are named and listed below:
- BioBricks
- BglBricks
One modification to the standard restriction enzyme method is the use of an enzyme that recognizes a non-palindromic sequence. This results in the ability to cut with a single restriction enzyme and still maintain directionality of the biological part.
- CpoI directional cloning
Other methods have been developed that utilize Type IIs restriction enzymes. These enzymes cut away from their recognition site. This strategy has the advantage over Type II enzymes in their reduction of a "scar" sequence and ability to generate combinatorial libraries.
Polymerase Chain Reaction
Polymerase Chain Reaction (PCR) is a fast and reliable way of making multiple copies of specific sequences of DNA. The PCR method typically allows for relatively simple and fast construction of genes by linking multiple shorter oligonucleotides together in a series of steps where the earlier PCR product acts as a primer that results in a longer DNA fragment that itself can act as a primer for subsequent rounds of PCR. An exception is TOPO TA Cloning (Invitrogen) [11][12]
, which utilizes the propert of Taq polymerase to leave a single A nucleotide overhang after the PCR reaction. In combination with an optimized vector with a T nucleotide overhang and a topoisomerase that recognizes the vector at the T overhang position, the insert and vector that are annealed at the T and A nucleotides are sealed into a circular vector efficiently.
Oligonucleotide precursors are linked together leaving gaps that must be filled in by DNA polymerase. In the most common method, polymerase cycling assembly (PCA) [13], DNA oligonucleotides are designed to be part of either the top or bottom of the final DNA duplex. Cycles of annealing and polymerase extension result in a growing DNA duplex built from smaller oligonucleotide fragments. A final round of PCR is done with constant primers that amplify the complete desired gene product. Initial oligo linkage is done for 20-30 cycles and full-length PCR is carried out for 20-30 additional cycles.
Oligos must be designed carefully to have similar melting temperatures and be free of interfering secondary structures (high GC content, hairpins, etc.)

Additional methods of gene assembly by PCR are listed below:
- Ligation Independent Cloning (LIC) [14][15][16]
- Uracil-Specific Excision Reagent (USER) (NEB) [17]
- Splice by Overlap Extension (SOE) PCR [18][19]
Recombination/Homology
- Cre/Lox P1 phage (BD Creator from Clontech)
- att lambda (Gateway Cloning from Life Technologies) [25][26]
- CPEC circular polymerase extension cloning [27]
- SLiCE (Seamless Ligation Cloning Extract) in vitro homologous recombination [28]
Gibson Assembly
In 2009, Daniel Gibson of J. Craig Venter's group reported a method for assembling hundreds of kilobases of DNA sequence using an isothermal enzymatic setup known today as "Gibson Assembly" [29][4][30][31]. Sub-fragments of the eventual assembly product are obtained as blunt-ended double-stranded DNA molecules several kilobases in length. They share several hundred bases of homology at their adjacent termini. T5 DNA Exonuclease chews back the double-stranded DNA molecule to reveal 3' single-stranded overhangs. Being heat-liable, T5 exonuclease is inactivated after several minutes at the reaction temperature. A mixture of thermostable ligase (Taq) and DNA polymerase (commonly Phusion) repair and assemble the individual fragments into larger molecules. This method has been used to assemble nonfunctional DNA molecules up to 900 kilobases in length, as well as the complete 16.3 kilobase mouse mitochondrial genome [32]. The size limit of molecules that can be assembled in this manner is currently limited to the cost of design and screening the final assembly products and the hundreds of kilobases of DNA that must be sequenced. An illustrative video of this process can be found on the NEB website. Of all the cloning techniques listed on this page, "Gibson" cloning is becoming one of the most dominant cloning strategies in synthetic biology labs. Its fast, flexible, relatively simple design has made it a favorite among synthetic biologists.

In Vivo
- Mating-assisted genetically integrated circuit (MAGIC)-bacterial mating [33]
- ET recombination [34]
- Transformation-associated recombination [35]
Moving Forward: Scaling Up Gene Synthesis
The methods below are not new methods and some are already being used commercially. What they offer for the future are methods for potentially scaling up production while decreasing costs. As mentioned before, cost and speed are major limitations preventing de novo DNA synthesis from becoming standard practice.
Solid-Phase Synthesis
An alternative gene synthesis technique using solid-phase synthesis of a DNA duplex has been created by several companies, including Blue Heron. In these methods, DNA oligonucleotides are synthesized as in other techniques, they are annealed into complementary short duplexes without overhanging ends, and these duplexes are chemically ligated on a solid phase such as a liquid chromatography column. The complete synthetic DNA duplex is then eluted and used for downstream processes, such as cloning or gene expression.
Multiplex/Microchip Synthesis
DNA oligonucleotides have become a commodity and the price per base has been steadily dropping. But there is still great expense involved in large-scale gene synthesis. For a several kilobase gene fragment, oligo cost alone can still exceed $1,000. To over come this hurdle, multiplex parallel gene synthesis technologies are under development, including inkjet DNA printers, photolithography and electrochemical parallel synthesis.
Microfluidic microchips are a common multiplexing solution. In the method of Tian et al. [36], oligonucleotides are synthesized while coupled to a microchip at their 3' end. The overall process is similar to traditional oligo synthesis, except that individual wells are photoactivated before addition of the next base onto the 5' end. After chemical cleavage of the complete oligo from the microchip, they are hybridized and purified using complementary "quality assurance" oligos bound to magnetic beads. This method eliminates oligos that have either incorporated extra bases or undergone deletions. Final yield from each synthesis well is only around 5 fmol of oligo, which is not sufficient for downstream applications. A final amplification step using common PCR primers to all oligos increases yields up to a million-fold. Oligos can then be assembled by overlap PCR-based methods such as polymerase assembly multiplexing, or PAM. This method has resulted in the synthesis, amplification and expression of a 14.5 kb, 21-gene 30S ribosomal protein operon from E. coli.
Minimizing Errors
One major roadblock remaining in synthetic gene synthesis is overcoming natural error rates in DNA synthesis chemistry. Phosphoramidite oligo synthesis currently has an error rate of 1 in ~160. This results in a high enough error rate in assembled genes that screening and sequencing are major cost bottlenecks, as is the repair or re-synthesis of mutated genes. While the development of cheap and high-throughput sequencing technologies will streamline the screening process, oligo synthesis error rates must also be reduced.
Currently, oligo pools are often screened against complementary reference libraries that can be reused. Oligos that do not completely base-pair with their designed sequence are either washed away in purification steps or are cleaved by DNA mismatch repair proteins due to their imperfect base-pairing. This has currently decreased error rates to 1 in 106, on the order of error rates in DNA polymerases.
Genome Synthesis
J. Craig Venter's team at JCVI have worked to carry gene synthesis to its most advanced level - the creation of a completely synthetic genome. Initially, the JCVI team synthesized a 582,970-base pair "minimized" genome of Mycoplasma genitalium using a stepwise assembly [29]. The genome was divided into 24 cassettes of about 24 kb each which were synthesized by commercial manufacturers. These overlapping cassettes were enzymatically stitched together into bacterial artificial chromosomes, which were attached in a stepwise fashion using a combination of restriction digestion and in vitro recombination. At the point where the molecules were "quarter genomes", they were transformed into yeast where in vivo recombination machinery assembled the full M. genitalium chromosome. This was only proof of synthesis, however, and not of a synthetic genome useful for supporting a living organism.
In 2010, the JCVI team reported the creation of a bacterium controlled by a fully synthetic genome[37]. The 1.08 mega-base pair genome of Mycoplasma mycoides was synthesized and transplanted into a M. capricolum recipient cell to create synthetic bacterium nicknamed "Synthia". The genome was divided into 1,078 1-kb cassettes, which were synthesized by Blue Heron. These cassettes were assembled in yeast using in vivo homologous recombination using a three-step hierarchy that built fragments of ~10,000 bp, ~100,000 bp and finally the complete 1.08 mega-bp genome. These were then isolated carefully from contaminating yeast DNA and transplanted into the recipient bacteria. A selectable marker placed in the synthetic genome demonstrated that bacteria that grew after transplantation were controlled by the synthetic genome. The same team demonstrated that a semi-synthetic genome could be assembled and transplanted using a combination of natural and synthetic DNA.
In 2011, a group from Johns Hopkins reported the first partially synthetic eukaryotic genome. They edited approximately 1% of the Saccharaomyces cerevisiae genome to eliminate introns, repetitive sequence and retrotransposons, replaced all TAG stop codons with TAA, and introduced recombigenic LoxPSym sites to aid in future large-scale manipulations of the genome [38].
Overall error rates in synthetic genome assembly were controlled by careful cassette synthesis and verification, as well as relying on high-fidelity in vivo recombination methods for hierarchical construction. Even low error rates can have huge effects, though. One single base deletion in the genome was in an essential gene, dnaA, and delayed progress for many weeks. Isolation, growth and manipulation of mega-base pair DNA molecules either in vivo or in vitro is a difficult roadblock as well.
Final assembly of large double-stranded DNA products into kilobase and megabase functional molecules, free of errors and with minimal cost remains a process bottleneck for gene and genome synthesis. In vitro recombination methods have been successfully used to assemble large DNA molecules on the order of tens of kilobases, but this method remains prone to errors and requires considerable homology be engineered into assembly fragments.
See Also
- Gibthon - Gibson assembly design program
- j5: A tool for designing DNA assembly with recombination from Nathan Hillson The manual has an excellent overview of recombination-based cloning strategies like SLIC and Gibson. [39]
- Review of Cloning techniques from ASU
References
- Khorana HG, Agarwal KL, Büchi H, Caruthers MH, Gupta NK, Kleppe K, Kumar A, Otsuka E, RajBhandary UL, Van de Sande JH, Sgaramella V, Terao T, Weber H, and Yamada T. Studies on polynucleotides. 103. Total synthesis of the structural gene for an alanine transfer ribonucleic acid from yeast. J Mol Biol. 1972 Dec 28;72(2):209-17. DOI:10.1016/0022-2836(72)90146-5 |
- Itakura K, Hirose T, Crea R, Riggs AD, Heyneker HL, Bolivar F, and Boyer HW. Expression in Escherichia coli of a chemically synthesized gene for the hormone somatostatin. Science. 1977 Dec 9;198(4321):1056-63. DOI:10.1126/science.412251 |
- Goeddel DV, Kleid DG, Bolivar F, Heyneker HL, Yansura DG, Crea R, Hirose T, Kraszewski A, Itakura K, and Riggs AD. Expression in Escherichia coli of chemically synthesized genes for human insulin. Proc Natl Acad Sci U S A. 1979 Jan;76(1):106-10. DOI:10.1073/pnas.76.1.106 |
- Gibson DG, Young L, Chuang RY, Venter JC, Hutchison CA 3rd, and Smith HO. Enzymatic assembly of DNA molecules up to several hundred kilobases. Nat Methods. 2009 May;6(5):343-5. DOI:10.1038/nmeth.1318 |
oligonucleotide assembly in vitro
- Hughes RA, Miklos AE, and Ellington AD. Gene synthesis: methods and applications. Methods Enzymol. 2011;498:277-309. DOI:10.1016/B978-0-12-385120-8.00012-7 |
Gene Synthesis Review
- Czar MJ, Anderson JC, Bader JS, and Peccoud J. Gene synthesis demystified. Trends Biotechnol. 2009 Feb;27(2):63-72. DOI:10.1016/j.tibtech.2008.10.007 |
review
- Engler C, Kandzia R, and Marillonnet S. A one pot, one step, precision cloning method with high throughput capability. PLoS One. 2008;3(11):e3647. DOI:10.1371/journal.pone.0003647 |
GoldenGate
- Weber E, Engler C, Gruetzner R, Werner S, and Marillonnet S. A modular cloning system for standardized assembly of multigene constructs. PLoS One. 2011 Feb 18;6(2):e16765. DOI:10.1371/journal.pone.0016765 |
MoClo
- Werner S, Engler C, Weber E, Gruetzner R, and Marillonnet S. Fast track assembly of multigene constructs using Golden Gate cloning and the MoClo system. Bioeng Bugs. 2012 Jan 1;3(1):38-43. DOI:10.4161/bbug.3.1.18223 |
MoClo
- Sarrion-Perdigones A, Falconi EE, Zandalinas SI, Juárez P, Fernández-del-Carmen A, Granell A, and Orzaez D. GoldenBraid: an iterative cloning system for standardized assembly of reusable genetic modules. PLoS One. 2011;6(7):e21622. DOI:10.1371/journal.pone.0021622 |
GoldenBraid
- Holton TA and Graham MW. A simple and efficient method for direct cloning of PCR products using ddT-tailed vectors. Nucleic Acids Res. 1991 Mar 11;19(5):1156. DOI:10.1093/nar/19.5.1156 |
TA cloning
- Shuman S. Recombination mediated by vaccinia virus DNA topoisomerase I in Escherichia coli is sequence specific. Proc Natl Acad Sci U S A. 1991 Nov 15;88(22):10104-8. DOI:10.1073/pnas.88.22.10104 |
TOPO
- Stemmer WP, Crameri A, Ha KD, Brennan TM, and Heyneker HL. Single-step assembly of a gene and entire plasmid from large numbers of oligodeoxyribonucleotides. Gene. 1995 Oct 16;164(1):49-53. DOI:10.1016/0378-1119(95)00511-4 |
PCA
- Aslanidis C and de Jong PJ. Ligation-independent cloning of PCR products (LIC-PCR). Nucleic Acids Res. 1990 Oct 25;18(20):6069-74. DOI:10.1093/nar/18.20.6069 |
LIC
- Aslanidis C, de Jong PJ, and Schmitz G. Minimal length requirement of the single-stranded tails for ligation-independent cloning (LIC) of PCR products. PCR Methods Appl. 1994 Dec;4(3):172-7. DOI:10.1101/gr.4.3.172 |
LIC
- Li C and Evans RM. Ligation independent cloning irrespective of restriction site compatibility. Nucleic Acids Res. 1997 Oct 15;25(20):4165-6. DOI:10.1093/nar/25.20.4165 |
LIC
- Geu-Flores F, Nour-Eldin HH, Nielsen MT, and Halkier BA. USER fusion: a rapid and efficient method for simultaneous fusion and cloning of multiple PCR products. Nucleic Acids Res. 2007;35(7):e55. DOI:10.1093/nar/gkm106 |
USER
- Higuchi R, Krummel B, and Saiki RK. A general method of in vitro preparation and specific mutagenesis of DNA fragments: study of protein and DNA interactions. Nucleic Acids Res. 1988 Aug 11;16(15):7351-67. DOI:10.1093/nar/16.15.7351 |
SOE-PCR
- Horton RM, Cai ZL, Ho SN, and Pease LR. Gene splicing by overlap extension: tailor-made genes using the polymerase chain reaction. Biotechniques. 1990 May;8(5):528-35.
SOEing
- Li MZ and Elledge SJ. Harnessing homologous recombination in vitro to generate recombinant DNA via SLIC. Nat Methods. 2007 Mar;4(3):251-6. DOI:10.1038/nmeth1010 |
SLIC
- Li MZ and Elledge SJ. SLIC: a method for sequence- and ligation-independent cloning. Methods Mol Biol. 2012;852:51-9. DOI:10.1007/978-1-61779-564-0_5 |
SLIC
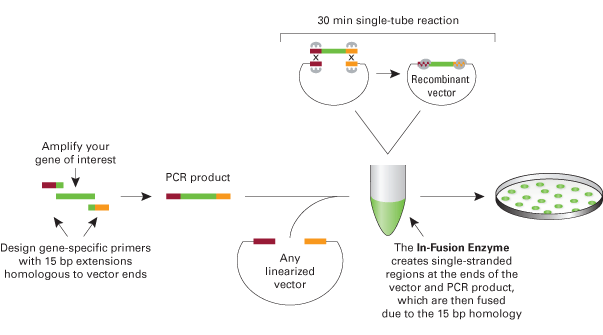
- Zhu B, Cai G, Hall EO, and Freeman GJ. In-fusion assembly: seamless engineering of multidomain fusion proteins, modular vectors, and mutations. Biotechniques. 2007 Sep;43(3):354-9. DOI:10.2144/000112536 |
in-fusion
- Benoit RM, Wilhelm RN, Scherer-Becker D, and Ostermeier C. An improved method for fast, robust, and seamless integration of DNA fragments into multiple plasmids. Protein Expr Purif. 2006 Jan;45(1):66-71. DOI:10.1016/j.pep.2005.09.022 |
in-fusion
- Sleight SC, Bartley BA, Lieviant JA, and Sauro HM. In-Fusion BioBrick assembly and re-engineering. Nucleic Acids Res. 2010 May;38(8):2624-36. DOI:10.1093/nar/gkq179 |
In-Fusion biobrick
- Hartley JL, Temple GF, and Brasch MA. DNA cloning using in vitro site-specific recombination. Genome Res. 2000 Nov;10(11):1788-95. DOI:10.1101/gr.143000 |
Gateway lambda Int
- Khalil AM, Julius JA, and Bachant J. One step construction of PCR mutagenized libraries for genetic analysis by recombination cloning. Nucleic Acids Res. 2007;35(16):e104. DOI:10.1093/nar/gkm583 |
Gateway lambda Cre
- Quan J and Tian J. Circular polymerase extension cloning of complex gene libraries and pathways. PLoS One. 2009 Jul 30;4(7):e6441. DOI:10.1371/journal.pone.0006441 |
CPEC
//T5 exonuclease recombination
- Zhang Y, Werling U, and Edelmann W. SLiCE: a novel bacterial cell extract-based DNA cloning method. Nucleic Acids Res. 2012 Apr;40(8):e55. DOI:10.1093/nar/gkr1288 |
SLiCe
- Gibson DG, Benders GA, Andrews-Pfannkoch C, Denisova EA, Baden-Tillson H, Zaveri J, Stockwell TB, Brownley A, Thomas DW, Algire MA, Merryman C, Young L, Noskov VN, Glass JI, Venter JC, Hutchison CA 3rd, and Smith HO. Complete chemical synthesis, assembly, and cloning of a Mycoplasma genitalium genome. Science. 2008 Feb 29;319(5867):1215-20. DOI:10.1126/science.1151721 |
Complete genome synthesis
- Gibson DG. Synthesis of DNA fragments in yeast by one-step assembly of overlapping oligonucleotides. Nucleic Acids Res. 2009 Nov;37(20):6984-90. DOI:10.1093/nar/gkp687 |
oligonucleotide assembly in yeast
- Gibson DG. Enzymatic assembly of overlapping DNA fragments. Methods Enzymol. 2011;498:349-61. DOI:10.1016/B978-0-12-385120-8.00015-2 |
MIE paper
- Gibson DG, Smith HO, Hutchison CA 3rd, Venter JC, and Merryman C. Chemical synthesis of the mouse mitochondrial genome. Nat Methods. 2010 Nov;7(11):901-3. DOI:10.1038/nmeth.1515 |
- Li MZ and Elledge SJ. MAGIC, an in vivo genetic method for the rapid construction of recombinant DNA molecules. Nat Genet. 2005 Mar;37(3):311-9. DOI:10.1038/ng1505 |
MAGIC, bacterial mating approach
- Zhang Y, Buchholz F, Muyrers JP, and Stewart AF. A new logic for DNA engineering using recombination in Escherichia coli. Nat Genet. 1998 Oct;20(2):123-8. DOI:10.1038/2417 |
ET recombination
- Larionov V, Kouprina N, Graves J, Chen XN, Korenberg JR, and Resnick MA. Specific cloning of human DNA as yeast artificial chromosomes by transformation-associated recombination. Proc Natl Acad Sci U S A. 1996 Jan 9;93(1):491-6. DOI:10.1073/pnas.93.1.491 |
Transformation-associated recombination (TAR) cloning
- Tian J, Gong H, Sheng N, Zhou X, Gulari E, Gao X, and Church G. Accurate multiplex gene synthesis from programmable DNA microchips. Nature. 2004 Dec 23;432(7020):1050-4. DOI:10.1038/nature03151 |
- Gibson DG, Glass JI, Lartigue C, Noskov VN, Chuang RY, Algire MA, Benders GA, Montague MG, Ma L, Moodie MM, Merryman C, Vashee S, Krishnakumar R, Assad-Garcia N, Andrews-Pfannkoch C, Denisova EA, Young L, Qi ZQ, Segall-Shapiro TH, Calvey CH, Parmar PP, Hutchison CA 3rd, Smith HO, and Venter JC. Creation of a bacterial cell controlled by a chemically synthesized genome. Science. 2010 Jul 2;329(5987):52-6. DOI:10.1126/science.1190719 |
genome replacement
- Dymond JS, Richardson SM, Coombes CE, Babatz T, Muller H, Annaluru N, Blake WJ, Schwerzmann JW, Dai J, Lindstrom DL, Boeke AC, Gottschling DE, Chandrasegaran S, Bader JS, and Boeke JD. Synthetic chromosome arms function in yeast and generate phenotypic diversity by design. Nature. 2011 Sep 14;477(7365):471-6. DOI:10.1038/nature10403 |
-
j5 DNA Assembly Design Automation Software doi: 10.1021/sb2000116