Biomod/2011/TeamJapan/Sendai/Computational design/Simulation: Difference between revisions
No edit summary |
No edit summary |
||
| Line 4: | Line 4: | ||
== About simulation == | == About simulation == | ||
We did 3D simulation of the molecular rolling robot over the DNA origami field using molecular dynamics. | We did 3D simulation of the molecular rolling robot over the DNA origami field using molecular dynamics. In this simulation, the robot represented as mass points, and moves by followed to Langevin equation. Hybridizing between robot legs and substrates on the field, and cleaving the substrates by legs, are described by potential shift. | ||
In this simulation, the robot | |||
Programs of our simulation were written in C language. | |||
== Model and Methods == | == Model and Methods == | ||
In this simulation, we used coarse-grained model (reference). As a coarse-grained model, representative points of structures were extracted, and structures were maintained by spring and string potentials. | |||
For example, see Fig.1, which represents extraction of corresponding points from real spider, and a scheme of potential to maintain structure. Streptavidine structures were maintained by spring-type potential which keeps length among points and weakly affect angles among points. Deoxyrybozyme legs were connected with string-type potential, which appears when points go beyond the determined length. On the spider simulation, blue points, green points, and yellow lines, represent mass points of the structure, top of spider legs, lines represents bonds, respectively. | |||
[[Image:Spider001.gif|600px|left|thumb|Fig.1 Spider in simulation]] | [[Image:Spider001.gif|600px|left|thumb|Fig.1 Spider in simulation]] | ||
<html><div style="clear:both;"></div></html> | <html><div style="clear:both;"></div></html> | ||
Each mass point is moving under influence of energy V. | Each mass point is moving under influence of energy V. The energy V is sum of potential energy for maintaining structure, and that for binding substrates on field. Potential of substrates is zero when legs are out of effective area (as cut-off), and changes by distance between legs and substrates when legs enter effective area. The force from differentiation of the substrate potential is proportional to the distance. | ||
Potential of substrates changes by distance of | |||
[[Image:potential.png|810px]] | [[Image:potential.png|810px]] | ||
| Line 46: | Line 25: | ||
The motion of each mass point is described by Langevin Equation. | The motion of each mass point is described by Langevin Equation. | ||
In this equation, force '''F''' | The motion of each mass point is described by Langevin Equation. In this equation, acceleration is determined by sum of force '''F''' from the differentiation of energy '''V''', viscosity resistance -β'''v''' , and white Gaussian random force η(t). White Gaussian force was obtained by box-muller methods (reference). Distribution of the obtained random force is shown in Fig.2. | ||
'''Langevin Equation''' | |||
[[Image:Langevin.png]] | [[Image:Langevin.png]] | ||
<html><div style="clear:both;"></div></html> | <html><div style="clear:both;"></div></html> | ||
| Line 60: | Line 35: | ||
<html><div style="clear:both;"></div></html> | <html><div style="clear:both;"></div></html> | ||
[[Image:Rforce.png|left|465px|thumb|Fig.2 Distribution of white Gaussian random force]] | |||
[[Image:Rforce.png|left|465px|thumb|Fig.2 | |||
<html><div style="clear:both;"></div></html> | <html><div style="clear:both;"></div></html> | ||
| Line 80: | Line 51: | ||
<td> | <td> | ||
In order to deduce parameters, we simulated movements of DNA spider robot in the DNA spider article (Lund et al, 2010). Red points and blue points represent uncleaved substrates and cleaved substrates, respectively. See the above movie, in which spider goes toward goal with cleaving substrates. | |||
In order to | |||
</td> | </td> | ||
</tr> | </tr> | ||
| Line 95: | Line 59: | ||
=== Simulation data === | === Simulation data === | ||
[[Image:Spider_data5.png|565px|thumb|right|Fig.4 (Left) Percentages of spiders that reached goal (without including the ones that left the field) (Right) Percentages of spiders which did not reach goal (which does not includes robots that was apart from field of DNA origami by brownian motion) These are resulted from 100 DNA spider simulation.]] | |||
Fig.4 describes how many spiders reached goal and how many did not at certain time. We tuned parameters on simulation to consist with the result in the figure 2G of DNA spider paper (Lund et al. 2010). On our simulation, the time for reaching the goal substrate was about 20 times longer than the time that DNAzyme cleaves substrate. The result matched very well to the time in the article (about 21 times longer). | |||
<html><div style="clear:both;"></div></html> | <html><div style="clear:both;"></div></html> | ||
| Line 138: | Line 77: | ||
</td> | </td> | ||
<td> | <td> | ||
We simulated whether our robot of triangular prism can reach goal of the field. Above videos shows our robot has a potential to reach goal. We supposed that our robot could reach the goal by only rotary motion, but however, our simulation result indicated that the triangular prism moves forward by corroboration manner of rotary motion with walking motion. This combined motion did not impede efficiency of moving forward. Anyway, simulation results reinforced us to consider our robots can reach goal. | |||
</td> | </td> | ||
</tr> | </tr> | ||
| Line 165: | Line 97: | ||
<html><div style="clear:both;"></div></html> | <html><div style="clear:both;"></div></html> | ||
To examine which is faster to reach goal, our triangular prism or DNA spider, simulation was done. Locations of substrates for the DNA spider were determined in above video by following the design of substrate in DNA spider paper. Locations of substrate field for triangular prism were followed to our field design. Since the body of the triangular prism robot is bigger than the spider body, we could reduce the number of substrates on the field. It can be seen from this video that the triangular prism robot reaches goal faster than the DNA spider. | |||
Since the body of the triangular prism robot is bigger than the spider body, we could reduce the number of substrates on the field. | |||
| Line 184: | Line 105: | ||
[[Image:Tri_data1.png|480px|thumb|right|Fig.6 Goal time and percentage of 1 leg type and 3 leg types robot]] | [[Image:Tri_data1.png|480px|thumb|right|Fig.6 Goal time and percentage of 1 leg type and 3 leg types robot]] | ||
To check the effect of the number of leg-types, speeds of triangular prism with 3 types of legs and with 1 type of legs were calculated by our simulation. The right part of Fig.6 shows the results. Goal time of both are very similar, but the percentage of triangle prism was different. 1 leg type robot has higher goal probability. Thus, we concluded that the 1 leg type robot is better. | |||
The right | |||
<html><div style="clear:both;"></div></html> | <html><div style="clear:both;"></div></html> | ||
Revision as of 02:01, 31 October 2011
<html>
<style rel="stylesheet" type="text/css">
.clear {clear:both;}
#verticalmenu {
/*this .CSS is inspired by http://www.javascriptkit.com/ */ font-family: "Comic Sans MS" , "Brush Script MT",serif, sans-serif, monospace, cursive, fantasy; list-style:none;}
- verticalmenu a:hover {
color: #aa1d1d; /* color when the click is over the main menu text-transform: uppercase; font-size: 10px; */
}
a:visited { color:#00a5ea; text-decoration: none }
.glossymenu, .glossymenu li ul{ list-style-type: none; margin: 0; padding: 0; width: 250px; /*WIDTH OF MAIN MENU ITEMS*/ border: 1px solid black; list-style:none; }
.glossymenu li{ position: relative; }
.glossymenu li a{ background: white url(http://openwetware.org/images/a/a7/Glossyback2.jpg) repeat-x bottom left; font: bold 17px Verdana, Helvetica, sans-serif; color: white; display: block; width: auto; padding: 10px 0; padding-left: 10px; text-decoration: none; }
.glossymenu li ul{ /*SUB MENU STYLE*/ position: absolute; width: 200px; /*WIDTH OF SUB MENU ITEMS*/
left: 0; top: 0; display: none; }
.glossymenu li ul li{ float: left; }
.glossymenu li ul a{ width: 190px; /*WIDTH OF SUB MENU ITEMS - 10px padding-left for A elements */ }
.glossymenu .arrowdiv{ position: absolute; right: 2px; background: transparent url(http://openwetware.org/images/6/69/Arrow.gif) no-repeat center right; }
.glossymenu li a:visited, .glossymenu li a:active{ color: white; }
.glossymenu li a:hover{ background-image: url(http://openwetware.org/images/5/50/Glossyback.jpg); }
/* Holly Hack for IE \*/
- html .glossymenu li { float: left; height: 1%; }
- html .glossymenu li a { height: 1%; }
/* End */
</style>
<script type="text/javascript">
/***********************************************
- CSS Vertical List Menu- by JavaScript Kit (www.javascriptkit.com)
- Menu interface credits: http://www.dynamicdrive.com/style/csslibrary/item/glossy-vertical-menu/
- This notice must stay intact for usage
- Visit JavaScript Kit at http://www.javascriptkit.com/ for this script and 100s more
- /
var menuids=new Array("verticalmenu") //Enter id(s) of UL menus, separated by commas var submenuoffset=-2 //Offset of submenus from main menu. Default is -2 pixels.
function createcssmenu(){ for (var i=0; i<menuids.length; i++){
var ultags=document.getElementById(menuids[i]).getElementsByTagName("ul")
for (var t=0; t<ultags.length; t++){
var spanref=document.createElement("span")
spanref.className="arrowdiv" spanref.innerHTML=" " ultags[t].parentNode.getElementsByTagName("a")[0].appendChild(spanref)
ultags[t].parentNode.onmouseover=function(){
this.getElementsByTagName("ul")[0].style.left=this.parentNode.offsetWidth+submenuoffset+"px"
this.getElementsByTagName("ul")[0].style.display="block"
}
ultags[t].parentNode.onmouseout=function(){
this.getElementsByTagName("ul")[0].style.display="none"
}
}
}
} if (window.addEventListener) window.addEventListener("load", createcssmenu, false) else if (window.attachEvent) window.attachEvent("onload", createcssmenu) </script>
<table border="0" align="center" vertical-align: middle;> <tr>
<td>
<ul id="verticalmenu" class="glossymenu"> <li><a href="http://openwetware.org/wiki/Biomod/2011/TeamJapan/Sendai">Home</a></li> <li><a href="http://openwetware.org/wiki/Biomod/2011/TeamJapan/Sendai/Strategy">Strategy</a></li> <li><a href="http://openwetware.org/wiki/Biomod/2011/TeamJapan/Sendai/Design">Design</a></li> <li><a href="#">Experiments</a>
<ul> <li><a href="http://openwetware.org/wiki/Biomod/2011/TeamJapan/Sendai/Results/Electrophoresis">Electrophoresis</a> <li><a href="http://openwetware.org/wiki/Biomod/2011/TeamJapan/Sendai/Results/Atomic_Force_Microscope">AFM</a> </ul>
</li> <li><a href="http://openwetware.org/wiki/Biomod/2011/TeamJapan/Sendai/Computational_design/Simulation" >Simulation</a></li> <li><a href="http://openwetware.org/wiki/Biomod/2011/TeamJapan/Sendai/Notes">Notes</a></li> <li><a href="http://openwetware.org/wiki/Biomod/2011/TeamJapan/Sendai/Team">Team</a></li> <li><a href="http://openwetware.org/wiki/Biomod/2011/TeamJapan/Sendai/Resources">Resources</a></li> <li><a href="http://openwetware.org/wiki/Biomod/2011/TeamJapan/Sendai/Sitemap">Sitemap</a></li>
</ul>
</td>
<td>
<img src="http://openwetware.org/images/0/07/3D_sendai.png" width="650"> </td> </tr> </table> </html> <html> <head> <style type="text/css">
- content {padding-left: 10px;width: 970px;}}
h3 {font-decoration: none;} h1.firstHeading {display: none; } </style> </head> </html>
<html> <style rel="stylesheet" type="text/css"> /*このスタイルシートの著作権はテンプレート工房TAKEにあります*/ /*ページのレイアウト用css*/
body{ background:#F5F5DC; /*壁色と壁紙設定*/ background-repeat:repeat;/*繰り返さない場合はno-repeatに変更*/ font:"メイリオ", "MS Pゴシック", Osaka, "ヒラギノ角ゴ Pro W3"; color: #333333; margin:0px; padding:0px; }
- contents{
width:900px;
margin:0 auto; background-color: #FFFFFF ;/*コンテンツ内の背景(サイズをぴったりにすること)*/ background-repeat:repeat-y; /*縦に繰り返し*/ border:solid 1px #666666;/*サイトに枠を付ける設定,色の変更可*/
position:relative;
font-size:80%;
}
/*ヘッダー部分の設定*/
- header{
background-image:url(http://openwetware.org/images/c/c4/TeamSendai-logo3.png) ;/*ヘーダー*/ background-repeat:repeat-x; /*縦に繰り返し*/ background-position:top right; height:140px; /*ヘーダーの高さ*/ }
- header p {
font-size: 26px;
color:#333333;
padding-top: 15px; padding-left: 20px; }
/*上部メニューボタンの設定*/
- navbar{
background-color:#FFFFFF;
width: 100%;
height:40px;
position:margine;
top:100px;
left:0px;
border-top:solid 1px #FFFFFF;
border-bottom:solid 1px #FFFFFF;
}
- navbar ul{
margin:0;
padding:0; list-style-type:none; font-family:Arial, Helvetica, sans-serif; font-size: 12px; line-height:40px; letter-spacing:2px; }
- navbar li{
background-color:#000099; /*上部メニューのボタンの背景*/
float:left; width:146px; /*メニューボタンの幅*/ text-align:center; padding:0; border-right:solid 1px #ffffff; }
- navbar ul a:hover{
background-color:#0033cc; /*メニューボタンにカーソルが来た時に背景*/
width:146px; /*メニューボタンの幅*/ }
- navbar a{
color:#ffffFF;/*メニューボタンの文字の色*/
display:block; }
- navbar a:hover{
color:#999999; /*メニューの文字がカーソルが来た時、この色に変わる*/ }
/*サイドメニューの設定*/
- side{
background-color:#ddffff;
width:220px;/*サイドの幅(変更するときはコンテンツ背景も変更すること)*/
position:margin; top:600px;/*上からの位置*/ left:12px; }
- side h3 {
font-size: 90%; border: double 3px #FFFFFF; color:#ffffff; text-align: center; background-color:#999999;
width:190px;
line-height: 30px; margin-top: 10px; margin-left: 5px; margin-bottom: 5px; }
- side h3 a {
color:#ffffff;
font-weight:normal; }
- side ul{
font-size:100%;
line-height:220%; /*サイドの文字と文字の行間設定*/ background-color: #ddffff; margin:0px; padding-left:10px; }
- side ul a:hover {
width:180px;
background-color: #99ffff; /*サイドのカーソルオーバー時の背景色*/ color: #999999; /*サイドのカーソルオーバー時の文字色*/ }
- side ul{
list-style-type:none;
padding-left:2px; }
- side li{
padding-left:15px; /*文字の左端からの位置*/ }
- side li a{
color:#333333;/*サイドの文字色*/
width:180px;
display:block;
}
- side .ad_list li{
background-image:none;
padding-left:0; }
/*右側メイン部分の設定*/
- main{
width:630px;
padding-top:15px;
margin-left:240px;
}
/*下部のフッター部分の設定*/
address{
font-size:80%;
font-style:normal;
text-align:center;
padding-top:5px;
}
address{
background-color:#000066;
color:#ffffff;
width:882px;
padding-bottom:10px; border:none; } address a{
color:#ff9999;
}
/*文字の設定*/ h1{ font-size:60%; letter-spacing: 2px; padding-left:10px; margin: 0px; }
h1 a{
color:#FFFFFF;
font-weight:normal; }
h2{
font-size:140%;
border-left: 10px solid #000066;
border-bottom:solid 1px #000099;/*文字の下に線を入れる設定*/
width:900px;
padding-left: 5px; color:#333333; margin-top: 15px; margin-bottom: 5px; }
h3{
font-size:120%;
border: solid 1px #111111;
color:#ffffff;
background-color:#4682B4 ; line-height: 30px; padding-left:10px; margin-top: 10px; margin-bottom: 1px; }
p{
font-size:90%;/*全体の文字サイズ*/
line-height:150%;/*全体で使う、文字と文字の行間*/
margin-left:5px;
}
p img{
float:left;
margin-top:5px; /*写真の左にスペースを空ける*/
margin-left:5px; /*写真の左にスペースを空ける*/
margin-right:10px; /:写真と文字の間隔*/ }
/*リンク文字の設定*/
a{
color:#000099;
text-decoration:none;
} a:hover { color: #FF0000;/*リンクの文字の上にマウスが来た時この色に変わる*/ text-decoration: none; }
- purple{
font-size:120%;
border: solid 1px #111111;
color:#ffffff;
background-color:#9370DB; line-height: 40px; padding-left:10px; margin-top: 10px; margin-bottom: 1px; }
h5{
font-size:120%;
border: solid 1px #111111;
color:#ffffff;
background-color:#FFA500; line-height: 30px; padding-left:10px; margin-top: 10px; margin-bottom: 1px; }
h6{
font-size:120%;
border: solid 1px #111111;
color:#ffffff;
background-color:#006400; line-height: 30px; padding-left:10px; margin-top: 10px; margin-bottom: 1px; }
- red{
font-size:120%;
border: solid 1px #111111;
color:#ffffff;
background-color:#DC143C; line-height: 40px; padding-left:10px; margin-top: 10px; margin-bottom: 1px; }
- blue{
font-size:120%;
border: solid 1px #111111;
color:#ffffff;
background-color:#191970; line-height: 40px; padding-left:10px; margin-top: 10px; margin-bottom: 1px; }
</style> </html>
About simulation
We did 3D simulation of the molecular rolling robot over the DNA origami field using molecular dynamics. In this simulation, the robot represented as mass points, and moves by followed to Langevin equation. Hybridizing between robot legs and substrates on the field, and cleaving the substrates by legs, are described by potential shift.
Programs of our simulation were written in C language.
Model and Methods
In this simulation, we used coarse-grained model (reference). As a coarse-grained model, representative points of structures were extracted, and structures were maintained by spring and string potentials.
For example, see Fig.1, which represents extraction of corresponding points from real spider, and a scheme of potential to maintain structure. Streptavidine structures were maintained by spring-type potential which keeps length among points and weakly affect angles among points. Deoxyrybozyme legs were connected with string-type potential, which appears when points go beyond the determined length. On the spider simulation, blue points, green points, and yellow lines, represent mass points of the structure, top of spider legs, lines represents bonds, respectively.

<html><div style="clear:both;"></div></html>
Each mass point is moving under influence of energy V. The energy V is sum of potential energy for maintaining structure, and that for binding substrates on field. Potential of substrates is zero when legs are out of effective area (as cut-off), and changes by distance between legs and substrates when legs enter effective area. The force from differentiation of the substrate potential is proportional to the distance.
 <html><div style="clear:both;"></div></html>
<html><div style="clear:both;"></div></html>
The motion of each mass point is described by Langevin Equation.
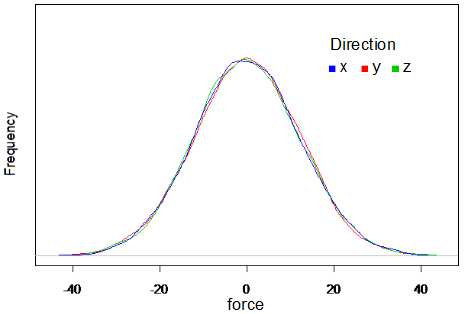
The motion of each mass point is described by Langevin Equation. In this equation, acceleration is determined by sum of force F from the differentiation of energy V, viscosity resistance -βv , and white Gaussian random force η(t). White Gaussian force was obtained by box-muller methods (reference). Distribution of the obtained random force is shown in Fig.2.
Langevin Equation
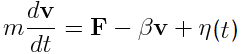 <html><div style="clear:both;"></div></html>
<html><div style="clear:both;"></div></html>
![]() <html><div style="clear:both;"></div></html>
<html><div style="clear:both;"></div></html>
<html><div style="clear:both;"></div></html>
Spider simulation

<html><div style="clear:both;"></div></html>
|
In order to deduce parameters, we simulated movements of DNA spider robot in the DNA spider article (Lund et al, 2010). Red points and blue points represent uncleaved substrates and cleaved substrates, respectively. See the above movie, in which spider goes toward goal with cleaving substrates. |
Simulation data

Fig.4 describes how many spiders reached goal and how many did not at certain time. We tuned parameters on simulation to consist with the result in the figure 2G of DNA spider paper (Lund et al. 2010). On our simulation, the time for reaching the goal substrate was about 20 times longer than the time that DNAzyme cleaves substrate. The result matched very well to the time in the article (about 21 times longer).
<html><div style="clear:both;"></div></html>
Simulation of the triangular prism robot
|
EmbedVideo received the bad id "z6_fWsgRH7A&border=1&color1=0x6699&color2=0x54abd6" for the service "youtube".
<html><div style="clear:both;"></div></html> |
We simulated whether our robot of triangular prism can reach goal of the field. Above videos shows our robot has a potential to reach goal. We supposed that our robot could reach the goal by only rotary motion, but however, our simulation result indicated that the triangular prism moves forward by corroboration manner of rotary motion with walking motion. This combined motion did not impede efficiency of moving forward. Anyway, simulation results reinforced us to consider our robots can reach goal.
|
Comparison between the spider and triangular prism robot

<html><div style="clear:both;"></div></html>
To examine which is faster to reach goal, our triangular prism or DNA spider, simulation was done. Locations of substrates for the DNA spider were determined in above video by following the design of substrate in DNA spider paper. Locations of substrate field for triangular prism were followed to our field design. Since the body of the triangular prism robot is bigger than the spider body, we could reduce the number of substrates on the field. It can be seen from this video that the triangular prism robot reaches goal faster than the DNA spider.
Data of triangular prism simulation

To check the effect of the number of leg-types, speeds of triangular prism with 3 types of legs and with 1 type of legs were calculated by our simulation. The right part of Fig.6 shows the results. Goal time of both are very similar, but the percentage of triangle prism was different. 1 leg type robot has higher goal probability. Thus, we concluded that the 1 leg type robot is better.
<html><div style="clear:both;"></div></html>
Other data
Cleavage rate and spider speed

To verify the effect of cleavage rate of substrate, we did spider simulation with changing cleavage time.
The following figure is the result we got from our simulation.
From this figure , we can see that percentage of spiders reached to the goal substrate increases and robot speed decreases if cleavage time increases.
This shows that increase of cleavage rate causes increase of spider speed and decrease of probability to reach the goal substrate.
<html><div style="clear:both;"></div></html>
Field design and spider speed
<html><div style="clear:both;"></div></html>

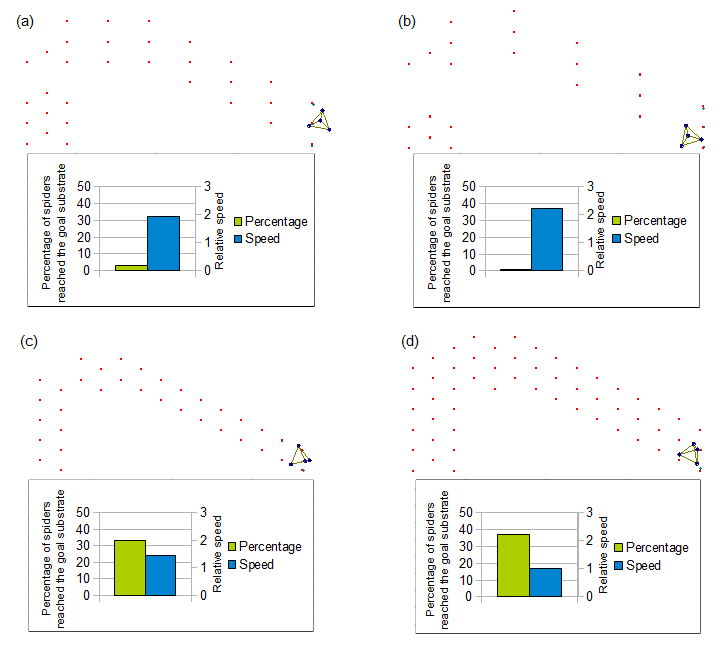
Above figure shows design of fields and moving data of spider over each field. Design of these fields are based on the field used in the spider article.(field of figure(d)) Substrates in field (a),(b),(c) is removed in different pattarn.
By looking each data from the viewpoint of the number of substrates, we got results in right figure. From this figure, we can see less substrates provides faster speed of robot. And we can also see the goal probability becomes higher if the number of substrates increases. But the goal probability seems to be strongly effected not only by the number but by its design too.
From these factors, we concluded that it is better to reduce substrates on the field as much as possible if we want to speed the robot.
<html><div style="clear:both;"></div></html>
Body size and leg size
<html><div style="clear:both;"></div></html>

In this section, whole size of both robots ,triangular prism and spider, are set up to be the same.
Design of triangular prism doesn't differ to the one we used above, while spider robot has longer legs.
Right figure shows moving data of each robot. From this figure, we can see the spider using long legs has more goal probability and goal time. This is thought to be due to the difference of body size, leg length and the number of legs. And faster speed in triangular prism robot can be seen in this simulation, too. If the robot has more legs, it can cleave more substrates in the same time, and then if the number of the substrates on the field is the same, robot has more legs will be faster. Thus, if the design of fields are the same, the robot has more legs is faster and this is why the triangular prism was faster in this simulation.
<html><div style="clear:both;"></div></html>