BioMicroCenter:Nextera: Difference between revisions
(New page: -- under construction-- rightNextera DNA sample preparation for Illumina sequencing from Epicentre technologies uses a modified transposase to simul...) |
(No difference)
|
Revision as of 13:35, 18 July 2011
-- under construction--

Nextera DNA sample preparation for Illumina sequencing from Epicentre technologies uses a modified transposase to simultaneously fragment and tag intact genomic DNA. A limited number of PCR steps are needed to create complete illumina libraries.
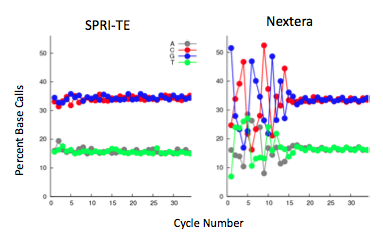
Nextera samples prepared in parallel with SPRI-te prepared illumina libraries provided similar quality sequencing data. DNA from the same Colobacter DNA sample were either sonicated or provided as intact DNA for preparation. The samples were multiplexed and sequenced on an illumina GAII, 40bp forward, in the same lane. The total genomic coverage for nextera and SPRI-te samples was exactly the same at 97.8% coverage of colobacter’s GC rich genome, although the complexity was greater in sonicated samples.

The nextera transposase does show mild GC insertion bias shown by the increase in percent of the first few bases. This, however, did not affect the sequencing coverage. The nextera prepared sample has very consistent coverage over the entire genome, while sonicated samples have a more variable coverage density.
We have used Nextera DNA sample preparation for multiple samples and have provided good quality data for all the samples. Below is an example of CNV study prepared with Nextera DNA preparation.