BME494 Sp2014 Dhatt: Difference between revisions
| (16 intermediate revisions by the same user not shown) | |||
| Line 40: | Line 40: | ||
<br><br> | |||
<!-- These are lines breaks for spacing purposes. You can add or delete these as needed --> | <!-- These are lines breaks for spacing purposes. You can add or delete these as needed --> | ||
| Line 49: | Line 49: | ||
<!-- Show a network/ circuit diagram of your device. Include a paragraph to explain how it works (e.g., how to switch the system from on to off and vice versa, and what happens to each component as the system switches between states) --> | <!-- Show a network/ circuit diagram of your device. Include a paragraph to explain how it works (e.g., how to switch the system from on to off and vice versa, and what happens to each component as the system switches between states) --> | ||
[[Image:BME494_Dhatt_AndGate.jpg|500px | [[Image:BME494_Dhatt_AndGate.jpg|500px|AND gate logic gene toggle switch. IPTG and low glucose levels conditions must be met in order for GFP production]] | ||
<br>AND gate logic gene toggle switch. IPTG and low glucose levels conditions must be met in order for GFP production.<br> | <br>AND gate logic gene toggle switch. IPTG and low glucose levels conditions must be met in order for GFP production.<br> | ||
[[Image:BME494_Dhatt_AndGateTable.jpg|500px | [[Image:BME494_Dhatt_AndGateTable.jpg|500px|Table describing that both inputs are needed in order to produce an output]] | ||
<br>Table describing that both inputs are needed in order to produce an output.<br> | <br>Table describing that both inputs are needed in order to produce an output.<br> | ||
<br><br> | |||
<!-- These are lines breaks for spacing purposes. You can add or delete these as needed --> | <!-- These are lines breaks for spacing purposes. You can add or delete these as needed --> | ||
| Line 68: | Line 68: | ||
[[Image:BME494_Dhatt_Assembly.jpg|500px|center]] | [[Image:BME494_Dhatt_Assembly.jpg|500px|center]] | ||
<!-- Show each DNA fragment in the order in which you propose to assemble them into a single plasmid. This can be a series of connected blocks. You do not have to show the plasmid as a circle. --> | <!-- Show each DNA fragment in the order in which you propose to assemble them into a single plasmid. This can be a series of connected blocks. You do not have to show the plasmid as a circle. --> | ||
| Line 76: | Line 76: | ||
<!-- List pre-existing parts/ Biobricks that you will use. Also list new parts that you will need to get from a natural source. BE SPECIFIC. --> | <!-- List pre-existing parts/ Biobricks that you will use. Also list new parts that you will need to get from a natural source. BE SPECIFIC. --> | ||
The following BioBrick parts, found on the iGEM registry website, will be used to build the new device: | The following BioBrick parts, found on the iGEM registry website, will be used to build the new device:<br> | ||
K418003 - composite Lac promoter inducible by IPTG | |||
K418003 - composite Lac promoter inducible by IPTG<br> | |||
Size: 1416 base pairs<br> | |||
LacI present: transcription inhibited<br> | |||
LacI absent: transcription promoted<br> | |||
B0030 – Strong RBS | LacI inhibited by IPTG<br> | ||
K259006 – Composite part made from green fluorescent protein (GFP) and double terminator | |||
B0030 – Strong RBS<br> | |||
Size: 15 base pairs<br> | |||
pSB1A3 – BioBrick assembly plasmid | |||
K259006 – Composite part made from green fluorescent protein (GFP) and double terminator<br> | |||
IPTG present: GFP produced<br> | |||
IPTG absent: GFP not produced, clear solution<br> | |||
Size: 823 base pairs<br> | |||
pSB1A3 – BioBrick assembly plasmid<br> | |||
Size:2155 base pairs<br> | |||
| Line 104: | Line 111: | ||
'''Primers'''<br> | '''Primers'''<br> | ||
The following primers will be used for the new device in order to introduce complementary overhangs and BsmBI cut sites. | The following primers will be used for the new device in order to introduce complementary overhangs and BsmBI cut sites.<br> | ||
Forward Primer | Forward Primer Vector: 5'-cacaccaCGTCTCaactagtagcggccgct<br> | ||
Reverse Primer | Reverse Primer Vector: 5'-cacaccaCGTCTCatctagatgcggccgcg<br> | ||
Forward Primer | Forward Primer Composite Lac Promoter: 5'-cacaccaCGTCTCatagattgacagctagctca<br> | ||
Reverse Primer | Reverse Primer Composite Lac Promoter: 5'-cacaccaCGTCTCatgtgtgtgctcagtatctt<br> | ||
Forward Primer RBS: 5'-cacaccaCGTCTCattaaagaggagaaa | Forward Primer RBS: 5'-cacaccaCGTCTCattaaagaggagaaa<br> | ||
Reverse Primer RBS: 5'-cacaccaCGTCTCaaattttctcctctttaat | Reverse Primer RBS: 5'-cacaccaCGTCTCaaattttctcctctttaat<br> | ||
Forward Primer GFP with terminator: 5'-cacaccaCGTCTCaatgcgtaaaggagaa | Forward Primer GFP with terminator: 5'-cacaccaCGTCTCaatgcgtaaaggagaa<br> | ||
Reverse Primer GFP with terminator: 5'-cacaccaCGTCTCatagtaaataataaaaaagc | Reverse Primer GFP with terminator: 5'-cacaccaCGTCTCatagtaaataataaaaaagc<br> | ||
Forward Primer for the mutation: 5’-agctgttgccGgtctcactgg | Forward Primer for the mutation: 5’-agctgttgccGgtctcactgg<br> | ||
Reverse Primer for the mutation: 5’-ccagtgagacCggcaacagct | Reverse Primer for the mutation: 5’-ccagtgagacCggcaacagct<br> | ||
| Line 157: | Line 162: | ||
4°C, ∞<br> | 4°C, ∞<br> | ||
<!-- Incorporate information from Presentation 2 --> | <!-- Incorporate information from Presentation 2 --> | ||
Digestion/Ligation Reaction Reagents:<br> | Digestion/Ligation Reaction Reagents:<br> | ||
| Line 210: | Line 216: | ||
<br><br><br> | |||
<!-- These are lines breaks for spacing purposes. You can add or delete these as needed --> | <!-- These are lines breaks for spacing purposes. You can add or delete these as needed --> | ||
| Line 219: | Line 225: | ||
I used a model of the natural Lac operon to learn how changing the parameter values changes the behavior of the system.<br> | I used a model of the natural Lac operon to learn how changing the parameter values changes the behavior of the system.<br> | ||
<!-- Continue this paragraph by explaining how you interacted with the MatLab model. Include two or more images showing different output curves that were generated when you altered the IPTG input concentration --> | <!-- Continue this paragraph by explaining how you interacted with the MatLab model. Include two or more images showing different output curves that were generated when you altered the IPTG input concentration --> | ||
[[Image:BME494_Dhatt_MRNA0.25.jpg|400px|]] | <br> | ||
[[Image:BME494_Dhatt_MRNA0.25.jpg|400px|]] | |||
[[Image:BME494_Dhatt_MRNA0.35.jpg|400px|]] | [[Image:BME494_Dhatt_MRNA0.35.jpg|400px|]] | ||
| Line 240: | Line 248: | ||
'''Differences''': <!-- Describe what aspects of your new device are not accounted for by the Lac operon model, and how this difference would require changes or additions to the Lac operon model. You may write this in general terms if you are not yet familiar with advanced biological modeling. Try your best! ---> | '''Differences''': <!-- Describe what aspects of your new device are not accounted for by the Lac operon model, and how this difference would require changes or additions to the Lac operon model. You may write this in general terms if you are not yet familiar with advanced biological modeling. Try your best! ---> | ||
The Lac operon model does not take into account glucose levels, which is an important component of the device and the and-logic gate. A model that incorporates both Lactose and glucose levels in order to test the device. | The Lac operon model does not take into account glucose levels, which is an important component of the device and the and-logic gate. A model that incorporates both Lactose and glucose levels would be essential in order to test the device. | ||
| Line 270: | Line 272: | ||
<br><br> | |||
==About the Designer== | ==About the Designer== | ||
| Line 276: | Line 278: | ||
<div style="color: #808080; background-color: #ffffff; width: 600px; padding: 5px"> | <div style="color: #808080; background-color: #ffffff; width: 600px; padding: 5px"> | ||
[[Image: | [[Image:BME494_Dhatt_profile.jpg |thumb|noframe|130px|left|'''Kulveen Dhatt''']] | ||
* My name is | * My name is Kulveen, and I am a senior majoring in Biomedical Engineering . I am taking BME 494 because synthetic biology sounded extremely interesting and it was a new topic of interest to me. An interesting fact about me is that I was born in India and moved here with my parents at a very young age.. | ||
</div> | </div> | ||
<br> | |||
==References== | ==References== | ||
<br> | |||
1. "American Diabetes Association". http://www.diabetes.org/<br> | |||
2. "Haynes:TypeIIS Assembly". http://openwetware.org/wiki/Haynes:TypeIIS_Assembly<br> | |||
3. Gardner T.S., Cantor C.Reverse, Collins J.J. 2000. Nature. Vol 409: 339-342.<br> | |||
4. iGEM Registry of Standard Biological Parts. http://parts.igem.org/Main_Page<br> | |||
Latest revision as of 17:13, 8 May 2014
My Profile Dr. Haynes OpenWetWare Previous Course Wiki Editing Help
Background & Proposed ApplicationBACKGROUND
The synthetic system modeled in the “Construction of a genetic toggle switch In Escherichia coli” uses two different states, an “on” state” and an “off” state (Gardner et all 2000). The classical system uses two repressor and two promoter pairs in which both repressors are inhibited by a different inducer, aTc and IPTG. The promoters were placed next to the repressor gene of the opposite pair. This system represents a bistable gene-regulatory network where only one promoter can be expressed at one time because the expression of one promoter repressed expression of the other promoter. The on-state of the cell was represented by placing a green fluorescent protein (GFP) transcription gene downstream of the on-state promoter. 
Globally, 227-285 million individuals have diabetes and about 90% of these individuals have Type II diabetes. In 2011, 1.4 million deaths occurred worldwide due to the result of diabetes making it the 8th leading cause of death. This number is estimated to double in the next 15 years, therefore, there is a need for a rapid diagnostic test.
Design of a New DeviceThe device will be designed as a diagnostic tool for diabetes and the functionality of the genetic switch replicates that of an “AND” logic gate. The device will require two conditions to be true in order for an output to be produced. One condition is that IPTG must be present in the device’s environment. When IPTG is present, it will bind to the LacI repressor and allow for transcription to continue. The other condition is that glucose levels in the device’s environment must be low. Glucose levels affect production of cAMP inversely; when glucose levels are high, cAMP production decreases and when glucose levels are low, cAMP production increases. cAMP binds to catabolite activator protein (CAP_ to form the CAP-cAMP complex. For the complex, cAMP must be present and glucose levels must be low. This complex is the required input of the device. In the natural lac operon, the CAP-cAMP complex leads to activation of gene expression from the lac operon. If glucose is present, cAMP levels will be low and the host will metabolize glucose if lactose is present.
Building the New DeviceSYNTHETIC DNA LAYOUT Type IIs Assembly was used to build this lac switch. Type IIs Assembly allows for parts to be assembled in one step. For Type IIs Assembly, forward and reverse primers are needed to be created and placed in the system in order to create sticky ends that can bind various parts together. To put the pieces together, PCR is implemented, which allows all the parts to be replicated thousands of times in order to produce a desired final product. Digestion and ligation is used during which BsmBI cuts the DNA fragments and creates complementary overhangs that anneal via base pairing. 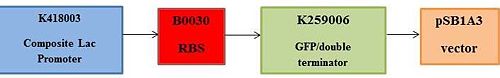
The following BioBrick parts, found on the iGEM registry website, will be used to build the new device: K418003 - composite Lac promoter inducible by IPTG
PCR Polymerase Chain Reaction (PCR) will be used for amplification of the DNA parts. PCR is the process of adding DNA, primers, nucleotides and DNA polymerase to a tube which produces system through assembly pot process after placed in a thermocycler and placed through many cycles. PCR consists of three specific temperature steps: denaturation, annealing, and elongation. Denaturation allows the DNA template to be heated to a specific temperature and yield single-stranded DNA molecules. The reaction temperature is then lowered for a short period of time in order to allow the complementary DNA primers to anneal to the single-stranded DNA template. The primers are then elongated by the DNA polymerase that is used. The DNA polymerase then allows for the cycle to be repeated multiple times resulting in thousands of copies of the desired fragments.
Forward Primer Vector: 5'-cacaccaCGTCTCaactagtagcggccgct Forward Primer Composite Lac Promoter: 5'-cacaccaCGTCTCatagattgacagctagctca Forward Primer RBS: 5'-cacaccaCGTCTCattaaagaggagaaa Forward Primer GFP with terminator: 5'-cacaccaCGTCTCaatgcgtaaaggagaa Forward Primer for the mutation: 5’-agctgttgccGgtctcactgg
Reagents
The PCR reagents used the following thermal cycling properties:
The Digestion/Ligation reagents used the following thermal cycling properties:
Testing the New DeviceLAC OPERON MODEL SIMULATION
RELATIONSHIP BETWEEN THE LAC MODEL AND MY NEW DESIGN
Differences: The Lac operon model does not take into account glucose levels, which is an important component of the device and the and-logic gate. A model that incorporates both Lactose and glucose levels would be essential in order to test the device.
Human Practices
DOES NOT SUPPORT AND UNDERSTANDS SUPPORTS BUT DOES NOT UNDERSTAND DOES NOT SUPPORT NOR UNDERSTANDS
About the Designer
References
|






