BME103:T130 Group 4 l2: Difference between revisions
Brent Russon (talk | contribs) (Undo revision 661001 by Brent Russon (Talk)) |
Brent Russon (talk | contribs) |
||
| (5 intermediate revisions by the same user not shown) | |||
| Line 159: | Line 159: | ||
==Research and Development== | ==Research and Development== | ||
<br> | |||
<!--- Bonus: explain how Bayesian statistics can be used to assess the reliability of your team's method. Just write the equation using variables that are relevant to your team's new test. You do not need actual numbers ---> | <!--- Bonus: explain how Bayesian statistics can be used to assess the reliability of your team's method. Just write the equation using variables that are relevant to your team's new test. You do not need actual numbers ---> | ||
'''Background on Disease Markers''' | '''Background on Disease Markers''' | ||
| Line 169: | Line 168: | ||
The associated gene change goes from a '''C'''GG (healthy gene sequence) to '''T'''GG (sequence associated with hemophilia). | The associated gene change goes from a '''C'''GG (healthy gene sequence) to '''T'''GG (sequence associated with hemophilia). | ||
The gene alteration leads to a mutated human protein. It goes from R[Arg] to W[Trp] | The gene alteration leads to a mutated human protein. It goes from R[Arg] to W[Trp] | ||
Hemophilia is a blood disorder. There are different types or degrees of hemophilia, but is essentially just different levels of blood not clotting properly. In it's most sever cases the body can have an incredibly hard time stopping a wound from bleeding because it will not clot to form a scab. Hemophilia is more likely to appear in males than in females because it is based in the X chromosome. | |||
| Line 186: | Line 187: | ||
<!--- Include an illustration that shows how your system's primers allow specific amplification of the disease-related SNP ---> | <!--- Include an illustration that shows how your system's primers allow specific amplification of the disease-related SNP ---> | ||
[[Image:Figure 1.png| | [[Image:Figure 1.png|800px|thumb|Figure 1]] | ||
[[Image:Figure 2.png| | [[Image:Figure 2.png|800px|thumb|Figure 2]] | ||
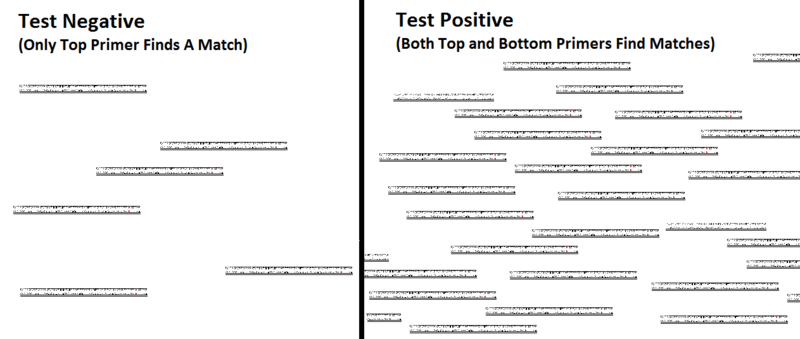
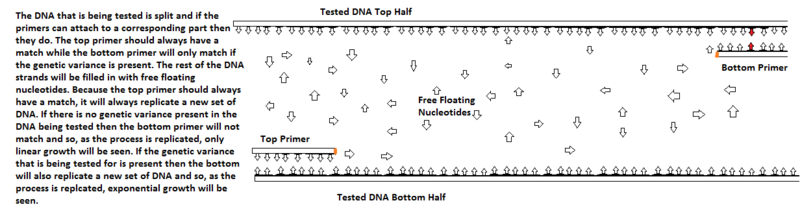
[[Image:Figure 3.png| | [[Image:Figure 3.png|800px|thumb|Figure 3: The DNA that is being tested is split and if the primers can attach to a corresponding part then they do. The top primer should always have a match while the bottom primer will only match if the genetic variance is present. The rest of the DNA strands will be filled in with free floating nucleotides. Because the top primer should always have a match, it will always replicate a new set of DNA. If there is no genetic variance present in the DNA being tested then the bottom primer will not match and so, as the process is replicated, only linear growth will be seen. If the genetic variance that is being tested for is present then the bottom will also replicate a new set of DNA and so, as the process is replicated, exponential growth will be seen.]] | ||
[[Image:Figure 4.png|800px|thumb|Figure 4]] | |||
<br> | |||
'''Bayesian Statistics''' | |||
<br> | |||
Because no test is completely accurate there are some useful statistics which can be calculated using Bayes rule after results are gathered. When looking at the probability that a person in a population has hemophilia, [ P(hc) ], the probability that if the test gives a positive result that you do in fact have hemophilia,[ P(T|hc) ], the probability that if the test gives a positive result that you do not have hemophilia, [ P(T|nc) ], and the probability that the test will give a positive result, [ P(T)], only three of them need to be known and then any other could be derived. This is demonstrated by the following equation: | |||
<br><br> | |||
(((100%-P(hc))x P(T|nc)) + ((P(hc) x P(T|hc)) = P(T) | |||
<br><br> | |||
<!-- ##### DO NOT edit below this line unless you know what you are doing. ##### --> | <!-- ##### DO NOT edit below this line unless you know what you are doing. ##### --> | ||
Latest revision as of 14:08, 29 November 2012
| Home People Lab Write-Up 1 Lab Write-Up 2 Lab Write-Up 3 Course Logistics For Instructors Photos Wiki Editing Help | |||||||||||||||||||||||||||||||||||||||||||||||||||
OUR TEAMLAB 2 WRITE-UPThermal Cycler EngineeringOur re-design is based upon the Open PCR system originally designed by Josh Perfetto and Tito Jankowski.
ProtocolsMaterials
Research and Development
Background on Disease Markers The marker that is being used is rs137852453. This SNP is associated with hemophilia. Hemophilia is a condition where someone's blood is not clotting properly. Data on this particular SNP variance can be seen here: http://www.ncbi.nlm.nih.gov/projects/SNP/snp_ref.cgi?rs=137852453 Hemophilia is a blood disorder. There are different types or degrees of hemophilia, but is essentially just different levels of blood not clotting properly. In it's most sever cases the body can have an incredibly hard time stopping a wound from bleeding because it will not clot to form a scab. Hemophilia is more likely to appear in males than in females because it is based in the X chromosome.
Bottom Primer (in reverse):
|
|||||||||||||||||||||||||||||||||||||||||||||||||||







