BME100 f2014:Group16 L5: Difference between revisions
From OpenWetWare
Jump to navigationJump to search
| (50 intermediate revisions by 5 users not shown) | |||
| Line 14: | Line 14: | ||
{| style="wikitable" width="700px" | {| style="wikitable" width="700px" | ||
|- valign="top" | |- valign="top" | ||
| [[Image: | | [[Image:MaebeOllie.jpg|100px|thumb|Name: Christopher Saar<br>Protocol editer<br> ]] | ||
| [[Image:royce.jpg|100px|thumb|Name: Gurpaul Sidhu<br>Role(s): Wiki editor, researcher<br>Picture Source:www. shadyrecords.com/wp-content/uploads/2013/01/royce.jpg]] | | [[Image:royce.jpg|100px|thumb|Name: Gurpaul Sidhu<br>Role(s): Wiki editor, researcher<br>Picture Source:www. shadyrecords.com/wp-content/uploads/2013/01/royce.jpg]] | ||
| [[Image:Bears kanye west 1280x1024 wallpaper wallpaperswa.com 75.jpg|100px|thumb|Name: Emily Angeles Mancinas<br>Role: assistant<br>Picture Source:http://wallpaperswa.com]] | | [[Image:Bears kanye west 1280x1024 wallpaper wallpaperswa.com 75.jpg|100px|thumb|Name: Emily Angeles Mancinas<br>Role: assistant<br>Picture Source:http://wallpaperswa.com]] | ||
| [[Image:moyai.jpg|100px|thumb|Name: Sheania Morgan<br>Roles: Conductor of experiment <br> image source: http://www.iemoji.com/view/emoji/814/places/moyai]] | | [[Image:moyai.jpg|100px|thumb|Name: Sheania Morgan<br>Roles: Conductor of experiment <br> image source: http://www.iemoji.com/view/emoji/814/places/moyai]] | ||
| [[Image: | | [[Image:Snoopy.jpg|100px|thumb|Name:Leslie Bernardino<br>Role: assistant/observer<br>source:www.fanpop.com ]] | ||
| [[Image:Francis.jpg|100px|thumb|Name: Romann Arizmendi <br> Assistant]] | | [[Image:Francis.jpg|100px|thumb|Name: Romann Arizmendi <br> Assistant]] | ||
|} | |} | ||
| Line 30: | Line 30: | ||
'''Smart Phone Camera Settings'''<br> | '''Smart Phone Camera Settings'''<br> | ||
<!-- The type of smart phone you used and how you adjusted the camera settings, if applicable. If you used more than one phone, make an additional list. --> | <!-- The type of smart phone you used and how you adjusted the camera settings, if applicable. If you used more than one phone, make an additional list. --> | ||
* Type of Smartphone: | * Type of Smartphone:'''iPhone 5s''' | ||
** Flash: '''NO Flash''' | ** Flash: '''NO Flash''' | ||
** ISO setting:''' | ** ISO setting:'''2000''' | ||
** White Balance: '''Auto''' | ** White Balance: '''Auto''' | ||
** Exposure: '''Zero''' | ** Exposure: '''Zero''' | ||
| Line 42: | Line 42: | ||
<!-- INSTRUCTIONS: Briefly describe how to set up your camera in front of the fluorimeter. Add a PHOTO of this set-up for bonus points. --> | <!-- INSTRUCTIONS: Briefly describe how to set up your camera in front of the fluorimeter. Add a PHOTO of this set-up for bonus points. --> | ||
* Distance between the smart phone cradle and drop = 6.5 cm | * Distance between the smart phone cradle and drop = '''6.5 cm''' | ||
'''Solutions Used for Calibration''' | '''Solutions Used for Calibration''' | ||
{| {{table}} border="3" cellspacing="3" width= | {| {{table}} border="3" cellspacing="3" width=700 | ||
|- | |- | ||
|align="center"| '''Init. Conc. 2X Calf Thymus DNA''' (micrograms/mL) ||align="center"|'''Vol. 2X DNA solution''' (micro-mL)||align="center"| '''Vol. SYBR GREEN 1 DYE''' (micro-mL) ||align="center"| '''Final DNA Concentration STBR GREEN 1 Solution''' (Micrograms/mL) | |align="center"| '''Init. Conc. 2X Calf Thymus DNA''' (micrograms/mL) ||align="center"|'''Vol. 2X DNA solution''' (micro-mL)||align="center"| '''Vol. SYBR GREEN 1 DYE''' (micro-mL) ||align="center"| '''Final DNA Concentration STBR GREEN 1 Solution''' (Micrograms/mL) | ||
|- | |- | ||
|align="center"| 5 ||align="center"| 80||align="center"| 80 ||align="center"| | |align="center"| 5 ||align="center"| 80||align="center"| 80 ||align="center"| 2.5 | ||
|- | |- | ||
|align="center"| 2 ||align="center"| 80 ||align="center"| 80 ||align="center"| | |align="center"| 2 ||align="center"| 80 ||align="center"| 80 ||align="center"| 1 | ||
|- | |- | ||
|align="center"| 1 ||align="center"| 80 ||align="center"| 80 ||align="center"| | |align="center"| 1 ||align="center"| 80 ||align="center"| 80 ||align="center"| 0.5 | ||
|- | |- | ||
|align="center"| 0.5 ||align="center"| 80 ||align="center"| 80 ||align="center"| | |align="center"| 0.5 ||align="center"| 80 ||align="center"| 80 ||align="center"| 0.25 | ||
|- | |- | ||
|align="center"| 0.25 ||align="center"| 80 ||align="center"| 80 ||align="center"| | |align="center"| 0.25 ||align="center"| 80 ||align="center"| 80 ||align="center"| 0.125 | ||
|- | |- | ||
|align="center"| 0 ||align="center"| 80 ||align="center"| 80 ||align="center"| | |align="center"| 0 ||align="center"| 80 ||align="center"| 80 ||align="center"| 0 | ||
|} | |} | ||
<!-- Add more rows and cells as needed. --> | <!-- Add more rows and cells as needed. --> | ||
| Line 67: | Line 67: | ||
'''Placing Samples onto the Fluorimeter''' | '''Placing Samples onto the Fluorimeter''' | ||
# | # Place fluorimeter on the table | ||
# | # Place slide in the fluorimeter with the smooth side down hydrophobic side up | ||
# | # Adjust slide to the light shines between the dots on the slide | ||
# '' | # Place '''80 micro-mL''' of <font color="green">'''SYBR green 1'''<font color="black"> on slide between the middle dots | ||
# '' | # Place '''80 micro-mL''' of calibration on the sphere you already placed to a '''160 micro-mL''' shpere | ||
# | # Adjust the shpere placement for the light and phone distance in the holder | ||
# | # Record Iphone distance from sample | ||
# | # Cover the assembly and make small adjustments | ||
# | # Take a photo on delay and cover | ||
# | # Send photo to be recorded for ''ImageJ'' analysis | ||
# '' | # Remove the '''160 micro-mL''' | ||
# | # Discard the slide | ||
# ''Repeat steps 1-12'' | # '''Repeat steps 1-12''' | ||
| Line 90: | Line 90: | ||
<!-- INSTRUCTIONS: (1) Show ONE image where you drew a circle around the droplet with the freehand tool for any sample with *no* DNA. (2) Show ONE image where you drew a circle around the droplet with the freehand tool for a sample *with* DNA (positive signal). -If you include more than two images, you will not receive any additional credit. --> | <!-- INSTRUCTIONS: (1) Show ONE image where you drew a circle around the droplet with the freehand tool for any sample with *no* DNA. (2) Show ONE image where you drew a circle around the droplet with the freehand tool for a sample *with* DNA (positive signal). -If you include more than two images, you will not receive any additional credit. --> | ||
<br> | <br> | ||
'''POSITIVE CONTROL IMAGE''' | {|{{table}}width=600 | ||
[[Image: | | align="center" style="background:#f0f0f0;"| '''POSITIVE CONTROL IMAGE'''|| align="center" style="background:#f0f0f0;"|'''NEGATIVE CONTROL IMAGE''' | ||
|- | |||
|[[Image:Pos234.jpg|200px|frame|center|Positive Sample]]||[[Image:Neg234.jpg|200px|frame|center|Negative Sample]] | |||
[[Image: | |- | ||
|} | |||
'''Image J Values for All Calibrator Samples''' | '''Image J Values for All Calibrator Samples''' | ||
| Line 102: | Line 102: | ||
'''EXCEL Table1 RAW DATA / Drop-Background''' | '''EXCEL Table1 RAW DATA / Drop-Background''' | ||
{|{{table}} width= | {|{{table}} width=550 | ||
| align="center" style="background:#f0f0f0;"|'''DNA Solution''' | | align="center" style="background:#f0f0f0;"|'''DNA Solution''' | ||
| align="center" style="background:#f0f0f0;"|'''Area''' | | align="center" style="background:#f0f0f0;"|'''Area''' | ||
| Line 110: | Line 110: | ||
| align="center" style="background:#f0f0f0;"|'''RAWINTDEN DROP-BACK''' | | align="center" style="background:#f0f0f0;"|'''RAWINTDEN DROP-BACK''' | ||
|- | |- | ||
|align="center"| '''0'''| | | align="center"|'''0'''||58604||61.036||3576976||175112||3401864 | ||
|- | |- | ||
|align="center"| '''0.25'''| | |align="center"| '''0.25'''||62324||88.828||5536096||175120||5360976 | ||
|- | |- | ||
|align="center"| '''0.5'''| | | align="center"|'''0.5'''||57291||108.156||6196361||165186||6031175 | ||
|- | |- | ||
|align="center"| '''1'''| | |align="center"| '''1'''||63052||152.1||9590219||189151||9401068 | ||
|- | |- | ||
|align="center"| '''2'''| | | align="center"|'''2'''||54628||195.326||10670272||140838||10529434 | ||
|- | |- | ||
|align="center"| '''5'''| | |align="center"|'''5'''||53200||222.41||11832237||144816||11687421 | ||
|- | |- | ||
|align="center"| '''Positive-1'''| | |align="center"| '''Positive-1'''||56188||199.791||11225837||138134||11087703 | ||
|- | |- | ||
|align="center"| '''Positive-2'''| | |align="center"| '''Positive-2'''||57469||202.95||11228718||140106||11088612 | ||
|- | |- | ||
|align="center"| '''Positive-3'''| | |align="center"| '''Positive-3'''||59569||203.44||11231472||142363||11089109 | ||
|- | |- | ||
|align="center"| '''Negative-1'''| | |align="center"| '''Negative-1'''||57504||76.644||4407348||132161||4275187 | ||
|- | |- | ||
|align="center"| '''Negative-2'''| | |align="center"| '''Negative-2'''||59222||73.25||4409466||134890||4274576 | ||
|- | |- | ||
|align="center"| '''Negative-3'''| | |align="center"| '''Negative-3'''||62109||72.05||4410288||137715||4272573 | ||
|- | |- | ||
|align="center"| ''' | |align="center"| '''P1 T1-1'''||65920||86.368||5693383||169276||5524107 | ||
|- | |- | ||
|align="center"| ''' | |align="center"| '''P1 T1-2'''||66209||83.35||5693737||169817||5523920 | ||
|- | |- | ||
|align="center"| ''' | |align="center"| '''P1 T1-3'''||69148||82.1||5694930||170726||5524204 | ||
|- | |- | ||
|align="center"| ''' | |align="center"| '''P1 T2-1'''||56864||73.951||4205157||142168||4062989 | ||
|- | |- | ||
|align="center"| ''' | |align="center"| '''P1 T2-2'''||59667||74.19||4205869||143593||4062276 | ||
|- | |- | ||
|align="center"| ''' | |align="center"| '''P1 T2-3'''||62540||77.27||4206231||145447||4060784 | ||
|- | |- | ||
|align="center"| ''' | |align="center"| '''P1 T3-1'''||62180||69.03||4292263||149505||4142758 | ||
|- | |- | ||
|align="center"| ''' | |align="center"| '''P1 T3-2'''||63625||67.76||4292430||151453||4140977 | ||
|- | |- | ||
|align="center"| ''' | |align="center"| '''P1 T3-3'''||64252||66.92||4292505||152708||4139797 | ||
|- | |- | ||
|align="center"| ''' | |align="center"| '''P2 T1-1'''||43144||230.772||9956425||177344||9779081 | ||
|- | |- | ||
|align="center"| ''' | |align="center"| '''P2 T1-2'''||43675||260.77||9958551||179669||9778882 | ||
|- | |- | ||
|align="center"| ''' | | align="center"|'''P2 T1-3'''||45197||238.14||9960640||180648||9779992 | ||
|- | |- | ||
|align="center"| ''' | |align="center"| '''P2 T2-1'''||48880||212.619||10392841||146922||10245919 | ||
|- | |- | ||
|align="center"| ''' | |align="center"| '''P2 T2-2'''||49197||205.44||10395760||148476||10247284 | ||
|- | |- | ||
|align="center"| ''' | |align="center"| '''P2 T2-3'''||50275||204.9||10395955||150674||10245281 | ||
|- | |- | ||
|align="center"| ''' | |align="center"| '''P2 T3-1'''||48880||212.619||10392841||121932||10270909 | ||
|- | |- | ||
|align="center"| ''' | |align="center"| '''P2 T3-2'''||49023||206.88||10392996||121968||10271028 | ||
|- | |- | ||
|align="center"| ''' | |align="center"| '''P2 T3-3'''||50014||205.17||10395265||122717||10272548 | ||
|} | |} | ||
<!-- Add more rows and cells as needed. --> | <!-- Add more rows and cells as needed. --> | ||
| Line 174: | Line 174: | ||
'''EXCEL Table 2 Edited to included background-subtractions''' | '''EXCEL Table 2 Edited to included background-subtractions''' | ||
{| {{table}} width= | {| {{table}} width=550 | ||
| align="center" style="background:#f0f0f0;"|'''Final DNA | | align="center" style="background:#f0f0f0;"|'''Final DNA Concentration in SYBR Green 1 Solution (ug/mL)''' | ||
| align="center" style="background:#f0f0f0;"|'''RAWINTDEN DROP-BACK''' | | align="center" style="background:#f0f0f0;"|'''RAWINTDEN DROP-BACK''' | ||
| align="center" style="background:#f0f0f0;"|''' | | align="center" style="background:#f0f0f0;"|''' ''' | ||
| align="center" style="background:#f0f0f0;"|''' | | align="center" style="background:#f0f0f0;"|''' ''' | ||
| align="center" style="background:#f0f0f0;"|''' | | align="center" style="background:#f0f0f0;"|''' ''' | ||
| align="center" style="background:#f0f0f0;"|'''Standard Deviation''' | | align="center" style="background:#f0f0f0;"|'''Standard Deviation''' | ||
|- | |- | ||
| ||'''1'''||'''2'''||'''3'''||'''MEAN'''|| | |align="center"|''' ''' ||align="center"|'''1'''||align="center"|'''2'''||align="center"|'''3'''||align="center"|'''MEAN'''|| | ||
|- | |- | ||
| 0||3401864|| | | align="center"|'''0'''||3401864||3401733||3403177||3402258||799 | ||
|- | |- | ||
| 0.25||5360976|| | |align="center"| '''0.25'''||5360976||5359811||5358887||5359891||1047 | ||
|- | |- | ||
| 0.5||6031175|| | |align="center"| '''0.5'''||6031175||6031020||6031474||6031223||231 | ||
|- | |- | ||
| 1||9401068|| | |align="center"| '''1'''||9401068||9399434||9399805||9400102||857 | ||
|- | |- | ||
| 2||10529434|| | |align="center"| '''2'''||10529434||10530655||10530916||10530335||791 | ||
|- | |- | ||
| 5||11687421|| | |align="center"| '''5'''||11687421||11687443||11689630||11688165||1269 | ||
|- | |- | ||
| Positive||11087703|| | |align="center"| '''Positive'''||11087703||11088156||11088712||11088190||505 | ||
|- | |- | ||
| Negative||4275187|| | |align="center"| '''Negative'''||4275187||4274013||4273439||4274213||891 | ||
|- | |- | ||
| | |align="center"| '''P1 T1'''||5524107||5524028||5522487||5523541||913 | ||
|- | |- | ||
| | |align="center"| '''P1 T2'''||4062989||4065331||4066390||4064903||1740 | ||
|- | |- | ||
| | |align="center"| '''P1 T3'''||4142758||4142815||4144110||4143228||765 | ||
|- | |- | ||
| | |align="center"| '''P2 T1'''||9779081||9778693||9778543||9778772||278 | ||
|- | |- | ||
| | |align="center"| '''P2 T2'''||10245919||10248449||10248278||10247549||1414 | ||
|- | |- | ||
| | |align="center"| '''P2 T3'''||10270909||10272563||10272912||10272128||1070 | ||
|} | |||
<br><br> | |||
'''INITIAL PCR PRODUCT CONCENTRATION = DNA x Total Dilution denominator''' | |||
{| {{table}} width=550 | |||
| align="center" style="background:#f0f0f0;"|'''PCR Prod Label''' | |||
| align="center" style="background:#f0f0f0;"|'''Mean RAWINTDEN Drop-Background''' | |||
| align="center" style="background:#f0f0f0;"|'''PCR Product Concentration (ug/mL)''' | |||
| align="center" style="background:#44AA22;"|'''Intial PCR Product Concentration (ug/mL)''' | |||
|- | |||
| align="center"|'''Positive'''||11088190||8.09||16.18 | |||
|- | |||
| align="center"|'''Negative'''||4274213||1.27||2.55 | |||
|- | |||
| align="center"|'''P1 T1'''||5523541||2.52||5.05 | |||
|- | |||
| align="center"|'''P1 T2'''||4064903||1.06||2.13 | |||
|- | |||
| align="center"|'''P1 T3'''||4143228||1.14||2.29 | |||
|- | |||
| align="center"|'''P2 T1'''||9778772||6.78||13.56 | |||
|- | |||
| align="center"|'''P2 T2'''||10247549||7.25||14.50 | |||
|- | |||
| align="center"|'''P2 T3'''||10272128||7.27||14.54 | |||
|- | |||
|} | |} | ||
<!-- Add more rows and cells as needed. --> | <!-- Add more rows and cells as needed. --> | ||
| Line 216: | Line 243: | ||
'''Calibration curve'''<br> | '''Calibration curve'''<br> | ||
<!-- INSTRUCTIONS: Place an image of your Excel plot with a line of best fit here. --> | <!-- INSTRUCTIONS: Place an image of your Excel plot with a line of best fit here. --> | ||
'''CONTROL CHART''' | |||
[[Image:DNACHART2-3.png|200px|frame|left|Control]] | |||
<br><br><br><br><br><br><br><br><br><br><br><br><br><br><br><br><br><br><br><br><br><br><br><br><br><br> | |||
'''PATIENT DNA''' | |||
[[Image:DNACHART1.png|200px|frame|left|Patient 1 & 2 Trials]] | |||
<br><br><br><br><br><br><br><br><br><br><br><br><br><br><br><br><br><br><br><br><br><br><br><br><br><br> | |||
'''PCR Results Summary''' | '''PCR Results Summary''' | ||
<!-- INSTRUCTIONS: You completed 8 PCR reactions and used the SYBR Green I staining and imaging technique to measure the amount of amplified DNA in each PCR reaction. You used a standard curve (based on known concentrations of calf thymus DNA) to convert INTDEN values into DNA concentration. Your positive control and negative control samples should be used as '''threshold''' values for determining whether an unknown (patient) sample is truly positive or negative. Replace the underscore with your claculated initial concentration values.--> | <!-- INSTRUCTIONS: You completed 8 PCR reactions and used the SYBR Green I staining and imaging technique to measure the amount of amplified DNA in each PCR reaction. You used a standard curve (based on known concentrations of calf thymus DNA) to convert INTDEN values into DNA concentration. Your positive control and negative control samples should be used as '''threshold''' values for determining whether an unknown (patient) sample is truly positive or negative. Replace the underscore with your claculated initial concentration values.--> | ||
* Our positive control PCR result was | * Our '''positive control PCR''' result was 11088190 μg/mL | ||
* Our negative control PCR result was | * Our '''negative control PCR''' result was 4274213 μg/mL | ||
<br> | |||
<u>Observed results</u> | <u>'''Observed results'''</u> | ||
<!-- INSTRUCTIONS: Replace the underscore with each patient ID. After the colon, write both a qualitative (what the images looked like) and a quantitative description (μg/mL) of what you observed --> | <!-- INSTRUCTIONS: Replace the underscore with each patient ID. After the colon, write both a qualitative (what the images looked like) and a quantitative description (μg/mL) of what you observed --> | ||
<br> | |||
{| {{table}} width=500 | |||
|align="center" style="background:#f0f0f0;"|'''PATIENT : 37465'''||align="center" style="background:#f0f0f0;"|'''PATIENT : 28404''' | |||
<u>Conclusions</u> | |- | ||
|[[Image:PATIENT1NEG.jpg|200px|frame|center|Patient 37436]]||[[Image:IMG_0116_2.jpg|200px|frame|center|Patient 28404]] | |||
|- | |||
|} | |||
{| {{table}} width=400 | |||
|align="center" style="background:#f0f0f0;"|'''NEGATIVE SAMPLE'''||align="center" style="background:#f0f0f0;"|'''PATIENT: 37465''' | |||
|- | |||
|align="center"|[[Image:NegSphere123.jpg|200px|frame|center|Negative Sample]]||align="center" |[[Image:PATIENT1NEG.jpg|200px|frame|center|Patient 37436]] | |||
|- | |||
|} | |||
{| {{table}} width=400 | |||
|align="center" style="background:#f0f0f0;"|'''POSITIVE SAMPLE'''||align="center" style="background:#f0f0f0;"|'''PATIENT: 28404''' | |||
|- | |||
|align="center"|[[Image:PosSphere123.jpg|200px|frame|center|Positive Sample]]||align="center" |[[Image:IMG_0116_2.jpg|200px|frame|center|Patient 28404]] | |||
|- | |||
|} | |||
<br><br><u>'''Conclusions'''</u><br> | |||
<!-- INSTRUCTIONS: Compare each patient's results to the positive control value and the negative control value. Draw a final conclusion for each patient (positive or negative) and explain why you made that conclusion. --> | <!-- INSTRUCTIONS: Compare each patient's results to the positive control value and the negative control value. Draw a final conclusion for each patient (positive or negative) and explain why you made that conclusion. --> | ||
{|{{table}} width=500 | |||
|- | |||
|Based upon the graph and the data, Patient 2 is positive and Patient 1 is negative. This is supported by the graph because Patient 2's data values lie above the line of best fit while Patient 1's data value lie below the regression line. | |||
|- | |||
|} | |||
<br><br> | <br><br> | ||
| Line 241: | Line 291: | ||
'''Background: About the Disease SNP''' | '''Background: About the Disease SNP''' | ||
<!-- INSTRUCTIONS: This content is from PCR Lab D. Write a summary, at least five sentences long, about the disease SNP in your own words. --> | <!-- INSTRUCTIONS: This content is from PCR Lab D. Write a summary, at least five sentences long, about the disease SNP in your own words. --> | ||
{|{{table}} | |||
|- | |||
| width=500 height=80|The disease is located on chromosome 21:34370656 on the KCNE2 gene. KCNE stands for potassium voltage-gated channel and joins with KCNH2 gene product, which is a pore forming protein, and alters its function. The mutation is a single nucleotide polymorphism and it is pathogenic in nature. It causes a Cardiovascular disease called Long QT syndrome. This mutation is found in Homo Sapiens by changing the genes that are responsible for encoding the ion channels present within the heart. The normal allele is TTC but it changes to CTC when a person has the disease. | |||
|- | |||
|} | |||
'''Primer Design and Testing''' | '''Primer Design and Testing''' | ||
<!-- INSTRUCTIONS: Write a short summary of the results of your primer test. Underneath your summary, include a screen capture of the results web page. You may crop the image so that it only includes the relevant information. --> | <!-- INSTRUCTIONS: Write a short summary of the results of your primer test. Underneath your summary, include a screen capture of the results web page. You may crop the image so that it only includes the relevant information. --> | ||
{|{{table}} | |||
|- | |||
|width=500 height=80|By inputing the non-disease forward primer and reverse forward primer the PCR database successfully matched the sequence meaning it is found in the healthy human genome. When the disease SNP was inputed with the single nucleotide change, the database could find no matches. | |||
|- | |||
|} | |||
[[Image:PCR Result1.png|200px|frame|left|Patient 28404]] | |||
<!-- Do not edit below this line --> | <!-- Do not edit below this line --> | ||
|} | |} | ||
Latest revision as of 15:52, 5 November 2014
| Home People Lab Write-Up 1 | Lab Write-Up 2 | Lab Write-Up 3 Lab Write-Up 4 | Lab Write-Up 5 | Lab Write-Up 6 Course Logistics For Instructors Photos Wiki Editing Help | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
OUR TEAM
LAB 5 WRITE-UPProcedureSmart Phone Camera Settings
Placing Samples onto the Fluorimeter
Data AnalysisRepresentative Images of Negative and Positive Samples
EXCEL Table1 RAW DATA / Drop-Background
EXCEL Table 2 Edited to included background-subtractions
INITIAL PCR PRODUCT CONCENTRATION = DNA x Total Dilution denominator
Calibration curve 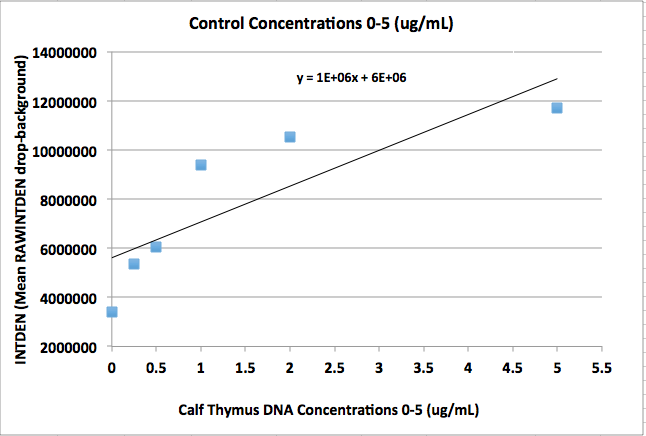

SNP Information & Primer DesignBackground: About the Disease SNP
Primer Design and Testing

| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||











