Axel:Schwekendiek Tutorial1
Basic cloning: How to construct new gene fragments?
This tutorial describes what we'll be doing in the lab. The procedures describe what you'll be doing and seeing during cloning. You'll have plenty of real-life experience with this procedure. First of all, let's examine an example:
Construction of GFP Biobrick 2.0 basic part PCR ca1123F/ca1123R on pSB1A2-I13522 (748 bp, BglII/XhoI/DpnI) Sub into pBca1102 (BglII/XhoI, 2159+697, L) Product is pBca1102-Bca1123 ---------------------------------------- ca1123F Forward BglII for eXtreme GFP basic part ctctgAGATCTatgcgtaaaggagaagaac ca1123R Reverse XhoI for eXtreme GFP basic part gcaaaCTCGAGttaGGATCCttatttgtatagttcatccatgc
Go ahead and download all the files and simulate the assembly on paper to confirm that it "works":
Media:JCAseq_pSB1A2-I13522.str Media:JCAseq_pBca1102.str Media:JCAseq_pBca1102-Bca1123.str
In this construction, we are using a plasmid pBca1102 I call "ORF expresser." This plasmid holds open reading frame basic parts, but it places a constitutive (meaning always on) promoter and ribosome binding site upstream of the Biobrick BglII site. The result of this is that the open reading frame will make the encoded protein when the plasmid is inserted into ‘’E. coli’’ cells. However, it is still a "basic part" in the sense that the ORF is still flanked by BamHI and BglII restriction sites. In this construction file, we start with the parent plasmid pBca1102 which has an RFP gene inside these restriction sites. Cells with pBca1102 are therefore red. When we digest with BglII/XhoI, we remove the RFP gene. We'll paste in the PCR-amplified GFP gene, and upon insertion of the new plasmid into E. coli, the cells should be green. So, that's the game here, now let's see how specifically to do it.
Step 1: The PCR
When you receive oligos from IDT, it comes as a lyophilized solid (ie, a dry powder). You must resuspend it in water to a final concentration of 100μM (μM is 10^-6 molar). Fortunately, IDT measures how many moles of DNA are present in the tube and writes it on the tube. Let's say the tube says it's 23.4nmoles of material. You would therefore add 234μL of water to the tube. I leave it to you to do the math and confirm that. To set up the PCR, you'll want to use 10μM stocks, so do a 10x dilution of the oligos. To achieve this, add 10μL of 100μM oligo to 90μL of water, and then mix it by tapping it on the bench. Now you can set up the PCR.
A general procedure for setting up a PCR reaction is described in the Standard PCR Setup:
Add the ingredients in order. In general, never add an enzyme to a reaction until after you've added the buffer. The reason for this is that proteins can denature if they are diluted into a solution that is not buffered. It doesn't matter when you add the oligos and template, though.
Now mix up the sample by tapping and place it in the thermocycler. Choose the temperature program based on the size of the fragment:
PCR product Size extension time Under 1kb 30 seconds 1kb to 2kb 60 sedonds 2kb to 4kb 4 minutes over 4kb 8 minutes
These programs principally differ in the length of time for each extension step during the cycles. This is necessary because longer PCR products take more time to polymerize. In general, it is OK to run a longer program than is necessary for the PCR product, but going shorter will cause failure. Sometimes doing longer extension times than are necessary will lead to aberrant products, so you should make an effort to use the right program. If you intend to clone the fragments after amplification, use 2 minutes extension. This gives you more A-overhangs. Alright, run the program.
Now you want to run an analytical gel to make sure you got the right product. There is another kind of gel you'll run later called a "preparative" gel. The distinction is that the analytical gel is run simply to examine the products. The preparative gel is done to separate DNA fragments, and bands are individually excised from the gel and purified. You will add 4μL of 6x loading buffer to the PCR reaction (assuming a standard 20 μL reaction volume), and then load 24 μL of material (all of it) to a well of the gel. In another lane, add 4μL of molecular weight marker. You'll use the marker to estimate the size of your PCR band. Below is an example of a PCR product.
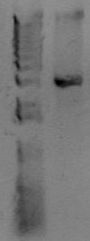

At the right of the PCR product is the known sizes present in the Hi-Lo molecular weight markers. The size of this PCR product is therefore somewhere around 3kb. After running this gel, estimate the size of your PCR product. Does it match the size you predicted for the construction file? Also, look at the quality of the band. Is it a single band or multiple bands? Is it tight or kindov fuzzy? If any of that fails, you'll need to troubleshoot. If it all looks good, let's move on to the cleanup reaction.
Step 2: Purify the PCR product
The general flow of the experiment goes 1) do the PCR, 2) clean up the PCR product, 3) digest the PCR product, 4) purify the digest. In parallel, you'll want to prepare the vector if it doesn't already exist. Then 5) set up the ligation reaction, and finally 6) transform the ligation.
You now need to purify the PCR product. We'll use the Fermentas kit for this. The binding buffer has a high concentration of guanidinium chloride. This salt forms a complex with nucleic acids which makes them bind to silica membranes. Also, it denatures proteins. So, adding binding buffer will destroy the polymerase present in the PCR. When you pass the mixture through the column, the DNA binds, but the denatured polymerase, dNTPs, buffer, and salts present in the mixture flow through. To carry out the procedure, just follow the instructions in the kit. And please do not forget to put the manual back into the box (the next group using it will thank you for that). You have to write the entire procedure down in your notebook, exactly as carried out.
The water re-dissolves the DNA which is now entirely pure. All that's left in your collection tube is DNA and water. This procedure achieves several things:
- It removes the polymerase which if left behind could either fill in or degrade the sticky ends you'll generate during the restriction digest
- It exchanges out the buffer because you'll want a different buffer for the restriction digest
- It removes most of the oligonucleotides and dNTPs left over from the PCR reaction, though they aren't much of an issue
Step 3: Set up the Restriction Digest
You now should have about 55-95 μL of DNA and solution, depending on if you eluted only once or twice. In order to increase the concentration, I recommend two elutions with 30μL that you keep separate. Check both for yield and use the highest concentrated sample for ligation (keep the other one as backup). If you want to increase the yield, you can use up to two time 50 μL. In any case, the first elution usually recovers less volume than water added, as some of the liquid tends to stay behind on the membrane. To assume only 25 μL recover if you elute with 30 μL, or 45 μL if yo elute with 50 μL. The second elution usually recovers all water added since the membrane is wet from the first elution. Buffers generally are made at 10x. So, set up the following reaction:
88.5μL DNA and water 10.0μL of Fermentas Fast Digest buffer 0.5μL BglII 0.5μL XhoI 0.5μL DpnI
The total amount of enzyme added can never exceed 10% of the final reaction volume. A larger fraction of total enzyme in the mix inhibits the reaction as the enzymes are stored in a glycerol solution. You are on the safe side here but watch this if you run smaller sample digests for checking. Tap the reaction on the bench to mix it thoroughly, give it a quick spin, then put it in the 37°C incubator for at least 15 minutes if you use Fermentas fast enzymes, and at least 1h if you use other enzymes.
Step 4: Clean up the Digest
After the digest has been completed, repeat the Fermentas cleaning procedure, elute two times with 30μL of water. This procedure will:
- Remove the leftover buffer
- Remove the restriction enzymes
- Partially remove the short DNA "tails" liberated by digestion
- Concentrate the DNA
Take a small aliquot (10% or μ3 L) to estimate the yield of recovery on an Agarose gel.
Step 5: Set up the ligation
Let's assume you already did the vector digest. Once purified, digested DNA will last for years in the freezer. So, you can save vector digests and use them over and over. In a later section, I'll describe how you do that digest, but for now, let's assume you already have done it. Set up the following reaction (in order):
6.5uL water 1uL 10x T4 DNA Ligase Buffer (in aliquots in freezer) 1uL vector Digest 1uL PCR product digest 0.5 uL T4 DNA Ligase
Make sure you are using T4 DNA ligase and not T4 DNA polymerase! Tap the reaction to mix, give it a quick spin, then put it on the bench (at room temp) to ligate for 30min. You can also leave the ligation over night (even days) at 4°C in the fridge.
To increase the yield of ligation product, I would strongly recommend to vary the vectoe:insert ratio. Play around with it and use for example 1:5 and 1:0.5 to see the effects. And do not forget to inlcude a negative control (only vector and no insert). Why is necessary?
Step 6: Do the transformation
Now we're ready to put the DNA into cells. There are 2 general ways to do this: heat shocking and electroporation. Both involve things called "competent cells". We make these in big batches and store them in the -80 freezer. We'll cover how to make them elsewhere. For now, just take one out of the -80 freezer and put it on ice to thaw out. There is 500 μL of cells in the aliquot. Add to it (on ice):
2-10 μL of your ligation mixture 50 μL competent cells
Do not forget to add a negative control (1. cells mixed with water instead of ligation product and 2. cells mixed with undigested vector). Why do you need both?
If you have more than 10 ligations in total, reduce the volume of cells you add to each of the ligations. Distribute the 500 μL of cells evenly among your reactions and do the no-DNA control last (just in case you don't have enough volume left).
Do not pipette up and down. Place the DNA first into a 15 ml tube and then add the 50 μL of cells on top of it. Gently move the tube to mix and place on ice immediately. Don't let it warm up--keep it on ice the whole time. Don't vortex it as cells are a little sensitive in this state.
Put your transformation mixtures on ice for 10 minutes, then do the heat shock for 90 sec. at 42°C (set up a water bath on time so that it will be ready when you need it). Then put it back on ice for 2 min (time isn't critical). Add 100 μL fresh and clear autoclaved SOC medium and mix.
Now you have to do different things based on the antibiotic marker you're going to use. If it is an ampicillin/carbenicillin resistance marker, you can plate immediately. This is because only growing bacteria are sensitive to the penicillin family of antibiotics. In contrast, all other antibiotics must be incubated at least 30 min. to give the cells time to express the marker. This is because most antibiotics directly inhibit translation. So, if you add the antibiotics immediately, you'll inhibit translation and the resistance protein never gets made. The cells will just die. So, instead, place the tubes into the shaker at 37°C for at least 30 minutes with gentle agitation (125 rpm). After that incubation, you can plate a the entire 100 μL of the reaction on the plates.
Now you're done for the day! Tomorrow you should have colonies, and you will go into the screening/characterization phase of the experiment.
Step 4b: Digesting the vector
In step 5 we set up ligations and assumed you already had a restriction digest of the vector (ie, plasmid pBca9145-Bca1089, pBca1102--the third line in your construction file). If you don't already have this in the freezer, you need to make it. You'll need a good quality miniprep (purified DNA) of your plasmid, then you'll set up the following:
3uL water 5uL miniprep 1uL Fermentas Fast Digest buffer 0.5uL restriction enzyme 1 0.5uL restriction enzyme 2
Often you'll want to scale the above numbers by a factor of 2 to 4. It all depends on how often you think you'll need this digest. Tap the mixture on the bench to mix, give it a quick spin, then put it at 37°C for 15 min. to one hour.
After incubating, add 2μL of 6x Loading Buffer (not marker!) load the entire mixture into one well of a 1% agarose gel. Put 4 μL of marker in another lane and run the gel. Make sure you wear gloves whenever working with agarose gels. The dye used to visualize DNA, ethidium bromide, is a carcinogen. Put the gel on a paper towel (take care that it does not slide off and drops on the floor) put it on the UV lamp. Be sure you are using the shield while doing this. Otherwise, you're basically staring into the sun--it's bad for your eyes. They can be seriously damaged. Now, look at the bands. Are they the size predicted by the construction file? Take a photograph and save it in your folder. And print a picture for each notebook.
Let's look at some gels:

A is an example of a "large" band from a plasmid digest. Usually, this is the sort of band you want. B is singly cut DNA. Often you'll predict 2 fragments from your digestion and will observe 3. If this is the case, look at the largest band--is it consistent with the full length of your plasmid? If so, it most likely is plasmid that only got cut once. The band you want is the middle band. In C you see an example of what I call a "smile" of the gel. Often some lanes of the gel run a little slower than others. If this happens, you can't just line up the bands with the marker to estimate the size. You have to mentally draw an arc through the gel. On this gel, all 4 lanes are the same plasmid, but as you can see they all look a little different. In D, you see a high molecular weight band in the lane. This can be one of two things. Sometimes, genomic DNA bleeds into your miniprep, and will look like a high molecular weight smear in the lane. Also, sometimes you get a "glycerol shadow", which is just a chemical phenomena--it's not DNA at all.
Cut out the appropriate band described in the construction file. Keep as little of the bordering agarose as possible, and transfer it to an microcentrifuge tube. Use the Fermentas gel extraction kit (not the gel purification kit). Really, what you need is a minimum of 3 gel volumes worth of ADB, but you'll need to estimate that (weigh out the tube before adding the piece and after). It won't hurt to add too much ADB, but you can add too little. Melt it at 55°C. Shake it occasionally, and keep heating until the whole thing is homogeneous (about 10 min.). Spin it through a column, wash twice with wash buffer, spin to dry, elute it in about same volume you initially started with (30-50 μL). But if you digested 10μL of miniprep, elute with 10μL of water only. If you did 5μL of miniprep, elute with 6μL of water--6 is the lowest volume you can elute with.