AVM BIOL368 Week 4: Difference between revisions
(→Part 1: add part 1 questions) |
(→Conclusion: finish conclusion) |
||
| (6 intermediate revisions by the same user not shown) | |||
| Line 50: | Line 50: | ||
==Data and Files== | ==Data and Files== | ||
*All files were saved to an external hard drive as well as on this page | *All files were saved to an external hard drive as well as on this page | ||
*[[Media:Visit_1_Subjects_1_thru_9_HIV.txt | HIV Sequences, Visit 1, Subjects 1-9]] | |||
*[[Media:Visit_1_Subjects_10_thru_15_HIV.txt | HIV Sequences, Visit 1, Subjects 10-15]] | |||
==Conclusion== | ==Conclusion== | ||
This exercise was done to determine if the HIV virus strains extracted from random test subjects originated from the same virus initially.Clones from subject 3 are the most genetically different of the subjects used. The tree shows that subjects 1 and 2 had very similar viruses, with some strains (subject 2; mutations 1 and 2, and subject 1; mutation 3) were even mores genetically similar. Subject 4 was slightly separated as well, but not as far away on the unrooted tree as subject 3. After following the protocol above and creating the unrooted tree during the exercise I understood that the extracted viruses did not stem from one original viral source. Overall, this project was helpful in helping to understand the significance of S values, theta, ClustalWdist, min and max differences and generally knowing how to understand similarities and differences between the sequences given. | |||
==Research Project== | ==Research Project== | ||
| Line 59: | Line 62: | ||
#Make a prediction (hypothesis) about the answer to your question before you begin your analysis. | #Make a prediction (hypothesis) about the answer to your question before you begin your analysis. | ||
#*There would be a shift in the subjects being grouped as moderate and rapid progressors. There is an overlap in virus number copies in Table 1 for subjects in both of these groups. | #*There would be a shift in the subjects being grouped as moderate and rapid progressors. There is an overlap in virus number copies in Table 1 for subjects in both of these groups. | ||
#Which subjects, visits, and clones will you use to answer your question? | #Which subjects, visits, and clones will you use to answer your question? Justify why you chose the subjects, visits, and clones you did. | ||
#*We could look at subjects 1, 3, 6 and 7 as they have very similar virus number copies. We could also choose subjects 12 and 13 to compare to those which have significantly lower virus number copies. | #*We could look at subjects 1, 3, 6 and 7 as they have very similar virus number copies. We could also choose subjects 12 and 13 to compare to those which have significantly lower virus number copies. For this specific question I think almost any of the subjects could be used and it may need to be adjusted once we actually start to work on the project. | ||
==Acknowledgments== | ==Acknowledgments== | ||
*Worked in collaboration with [[User:Courtney L. Merriam| Courtney L. Merriam]] in class. | *Worked in collaboration with [[User:Courtney L. Merriam| Courtney L. Merriam]] in class. Courtney and I agreed on the topic and details of our research in class, and wrote the section titled "Defining Our Research Project" together. | ||
*Template followed from week 4 assignment [http://openwetware.org/wiki/BIOL368/F16:Week_4 BIOL368/F16] | *Template followed from week 4 assignment [http://openwetware.org/wiki/BIOL368/F16:Week_4 BIOL368/F16] | ||
*Help from our professor, [[User:Kam D. Dahlquist|Dr. Dahlquist]] in class. | *Help from our professor, [[User:Kam D. Dahlquist|Dr. Dahlquist]] in class. | ||
| Line 74: | Line 75: | ||
==References== | ==References== | ||
* Donovan S and Weisstein AE (2003) Exploring HIV Evolution: An Opportunity for Research. In Jungck JR, Fass MR, and Stanley ED, eds. ''Microbes Count!'' West Chester, Pennsylvania: Keystone Digital Press. | |||
* Markham, R.B., Wang, W.C., Weisstein, A.E., Wang, Z., Munoz, A., Templeton, A., Margolick, J., Vlahov, D., Quinn, T., Farzadegan, H., & Yu, X.F. (1998). Patterns of HIV-1 evolution in individuals with differing rates of CD4 T cell decline. ''Proc Natl Acad Sci U S A. 95'', 12568-12573. doi: 10.1073/pnas.95.21.12568 | |||
** [http://www.pnas.org/content/95/21/12568.long Publisher Full text] | |||
**[http://www.pnas.org/content/95/21/12568.full.pdf+html PDF]. | |||
* Vlahov, D., Anthony, J.C., Munoz, A., Margolick, J., Nelson, K.E., Celentano, D.D., Solomon, L., Polk, B.F. (1991). The ALIVE study, a longitudinal study of HIV-1 infection in intravenous drug users: description of methods and characteristics of participants. ''NIDA Res Monogr 109'', 75-100. | |||
** [http://www.ncbi.nlm.nih.gov/pubmed/1661376 PubMed] | |||
** [http://www.nida.nih.gov/pdf/monographs/109.pdf PDF]. | |||
*[[BIOL368/F16:Week 4 | Week 4 Assignment Page]] | |||
<!-- copied references from group page --!> | |||
{{Template: Avery_Vernon-Moore}} | {{Template: Avery_Vernon-Moore}} | ||
Revision as of 22:07, 25 September 2016
Purpose
- To continue with our class research and journal projects for the Markham et al. paper
- Continue to learn about sequencing and the similarities and differences
Methods and Results
Part 1
- Upload the “visit_1_S1_S9.txt” and “visit_1_S10_S15.txt” files from the BIOL368/F16 Week 4 page into the Biology Workbench
- Generate a multiple sequence alignment using ClustlW
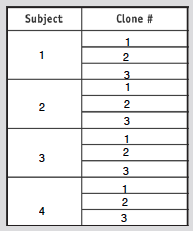
- Choose 3 clones form 4 different subjects (as shown in table below)
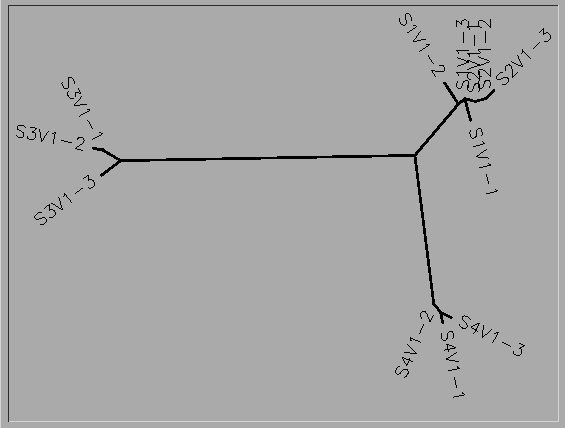
- Photo of tree shown below
Questions
- Do the clones from each subject cluster together?
- Yes
- Do some subjects’ clones show more diversity than others?
- Subject 1, strain 1 is more diverse than strains 2 and 3. Subject 2 strain 3 is more diverse than stains 1 and 2. Subject 3 strain 3 is more diverse than strains 1 and 2. Subject 4 strain 2 is more diverse than strains 1 and 3.
- Do some of the subjects cluster together?
- Yes. Subjects 1 and 2 are clustered.
- Write a brief description of the tree and how to interpret the clustering pattern with respect to the similarities and potential evolutionary relationships between subjects’ HIV sequences.
- The unrooted tree shows the similarities of the 4 subjects and clones. This tree shows that Subject 1 and 2 have very closely related viruses, and within this virus, mutations 1 and 2 of subject 2 are most closely related to mutation 3 of subject 1. All of the mutations of subject 4's virus are most closely related to each other and the mutations of subject 3 are most closely related to each other.
Part 2
- Select a sequence from last weeks assignment and align all of the clones
- Calculate S from the alignment
- S is the number of positions where there is at least one nucleotide difference across the clones
- Save the alignment for later use
- Run an analysis for two more subjects
- Sequence differences for each subject shown at right



- (Math Is Fun was used to calculate theta)
- Within the Clustdist tool generate a distance matrix for one of your alignments
- Choose the highest and lowest pairwise scores and find the % difference
- Calculate theta for each subject
- Repeat these steps for each subject
- Table below shows results from these analysis
- Create a new alignment with all the sequences from 2 different subjects
- Use Clustdist to create a pairwise distance matrix
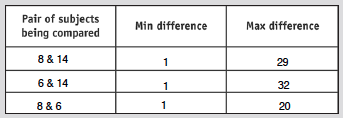
- Find all the min and max differences
- Results shown in table below
Data and Files
- All files were saved to an external hard drive as well as on this page
- HIV Sequences, Visit 1, Subjects 1-9
- HIV Sequences, Visit 1, Subjects 10-15
Conclusion
This exercise was done to determine if the HIV virus strains extracted from random test subjects originated from the same virus initially.Clones from subject 3 are the most genetically different of the subjects used. The tree shows that subjects 1 and 2 had very similar viruses, with some strains (subject 2; mutations 1 and 2, and subject 1; mutation 3) were even mores genetically similar. Subject 4 was slightly separated as well, but not as far away on the unrooted tree as subject 3. After following the protocol above and creating the unrooted tree during the exercise I understood that the extracted viruses did not stem from one original viral source. Overall, this project was helpful in helping to understand the significance of S values, theta, ClustalWdist, min and max differences and generally knowing how to understand similarities and differences between the sequences given.
Research Project
- What is your question?
- If HIV-1 virus evolution patterns are based on virus copy number rather than CD4 T-cell decline, how would this change the subjects classification as non-progressors, moderate progressors, and rapid progress ors? Is virus copy number correlated with diversity and divergence?
- Make a prediction (hypothesis) about the answer to your question before you begin your analysis.
- There would be a shift in the subjects being grouped as moderate and rapid progressors. There is an overlap in virus number copies in Table 1 for subjects in both of these groups.
- Which subjects, visits, and clones will you use to answer your question? Justify why you chose the subjects, visits, and clones you did.
- We could look at subjects 1, 3, 6 and 7 as they have very similar virus number copies. We could also choose subjects 12 and 13 to compare to those which have significantly lower virus number copies. For this specific question I think almost any of the subjects could be used and it may need to be adjusted once we actually start to work on the project.
Acknowledgments
- Worked in collaboration with Courtney L. Merriam in class. Courtney and I agreed on the topic and details of our research in class, and wrote the section titled "Defining Our Research Project" together.
- Template followed from week 4 assignment BIOL368/F16
- Help from our professor, Dr. Dahlquist in class.
- Note: While I worked with the people noted above, this individual journal entry was completed by me and not copied from another source.
Avery Vernon-Moore 14:42, 21 September 2016 (EDT)
References
- Donovan S and Weisstein AE (2003) Exploring HIV Evolution: An Opportunity for Research. In Jungck JR, Fass MR, and Stanley ED, eds. Microbes Count! West Chester, Pennsylvania: Keystone Digital Press.
- Markham, R.B., Wang, W.C., Weisstein, A.E., Wang, Z., Munoz, A., Templeton, A., Margolick, J., Vlahov, D., Quinn, T., Farzadegan, H., & Yu, X.F. (1998). Patterns of HIV-1 evolution in individuals with differing rates of CD4 T cell decline. Proc Natl Acad Sci U S A. 95, 12568-12573. doi: 10.1073/pnas.95.21.12568
- Vlahov, D., Anthony, J.C., Munoz, A., Margolick, J., Nelson, K.E., Celentano, D.D., Solomon, L., Polk, B.F. (1991). The ALIVE study, a longitudinal study of HIV-1 infection in intravenous drug users: description of methods and characteristics of participants. NIDA Res Monogr 109, 75-100.
- Week 4 Assignment Page