20.109(S13):Bacterial amplification of DNA (Day3): Difference between revisions
(New page: {{Template:20.109(S13)}} <div style="padding: 10px; width: 640px; border: 5px solid #99FF66;"> ==Introduction== [[Image:Be109transformation.jpg|thumb|right|225px|'''Bacterial transform...) |
(No difference)
|
Revision as of 17:18, 28 January 2013
Introduction
Assuming all went well, your reaction tubes from last time contain mutagenized DNA that encodes mutant inverse pericam. However, the desired DNA plasmid is likely present at a low concentration, and moreover it is nicked rather than in intact circular form. What we would like to do now is repair and further amplify only the mutagenized product. Thankfully, we have E. coli bacteria to do this for us quite efficiently!
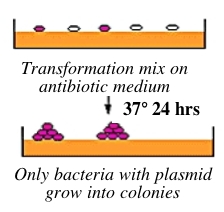
Bacteria can take up foreign DNA in a process called transformation, during which a single plasmid enters a bacterium and, once inside, replicates and expresses the genes it encodes. Most bacteria do not exist in a transformation-ready state, but can be made permeable to foreign DNA by chemical treatment or other means. Cells that are capable of transformation are referred to as competent. Competent cells are extremely fragile and should be handled gently, i.e., kept cold and not vortexed. Bacterial transformation is efficient enough for most lab purposes, resulting in as many as 109 transformed cells per microgram of DNA, but even with highly competent cells only 1 DNA molecule in about 10,000 is successfully transformed. Thus we need a way to identify transformed cells, which is usually accomplished with antiobiotics. For example, the plasmid carrying inverse pericam (called pRSET) also carries a gene that leads to ampicillin-resistance. Consequently, a transformed bacterium will grow on ampicillin-containing agar medium, while untransformed cells will die before they can form a colony (see figure above right). Given the low concentration and nicked structure of your DNA to begin with, you should perform your transformations today with great care.
Before setting up transformations, you will test your mutagenized DNA for the presence and approximate concentration of product via gel electrophoresis. Because the product is several Kbp long, a standard 1% agarose gel will serve us just fine. The long mutant plasmid DNA should be separated from the short digested fragments of parental DNA and thus can be identified. Note that the parental plasmid is originally present at a concentration too low to detect on a gel, and even the amplified mutant may show up as only a faint band. If you do not see any band at the expected size of the mutant plasmid, you might increase the amount of DNA used during the transformation procedure at the end of lab.
Protocols
Part 1: Agarose gel electrophoresis
Using a 1% agarose gel prepared by the teaching faculty, you will each run your digested reaction mixture. Most of you will also load either a reference lane containing standards of known molecular weight or a somewhat concentrated sample of wild-type plasmid. Divide up that responsibility however you see fit. Recall that you should always handle all gels and gel equipment with nitrile gloves.
- In an eppendorf tube, combine 10 μL of of your DpnI-digested product with 2.5 μL of loading dye. Save the rest of the digested DNA, keeping it on ice.
- Remember to flick and quick-spin your sample, or pipet up and down to mix.
- Recall that loading dye contains xylene cyanol as a tracking dye and glycerol to help the samples sink into the well.
- Load the gel in the order shown in the table below, 11 μL per SDM sample and 10 μL per ladder lane.
- We will use a 14-lane rather than a 10-lane gel, to concentrate the DNA into a smaller area.
- To load, lower your sample-containing tip below the surface of the buffer and directly over the well. Expel your sample into the well. Do not release the pipet plunger until after you have removed the tip from the gel box.
- Once all the samples have been loaded, the teaching faculty will attach the gel box to the power supply and run the gel at 100 V for 45 minutes.
| Lane | Sample | Lane | Sample |
|---|---|---|---|
| 1 | Red group | 8 | Parent IPC |
| 2 | Orange group | 9 | Blue group |
| 3 | DNA Ladder | 10 | Pink group |
| 4 | Parent IPC | 11 | DNA Ladder |
| 5 | Yellow group | 12 | Parent IPC |
| 6 | Green group | 13 | Purple group |
| 7 | DNA Ladder | 14 | BLANK |
While the gel runs, you can work on Parts 3 and (optional) 5, label the tubes you will need in Part 4, work on your notebooks, start the FNT assignment, etc. Be sure to pre-chill your 14 mL tubes on ice for at least a few min before adding competent cells to them.
Part 2: Gel analysis
- The teaching faculty will photograph and post a digital image of the gel.
- In the following analysis, you will need the information for the 1 Kbp ladder you used, which is available at this link.
- First, see if you got a band at the expected size of the pRSET plasmid with an inverse pericam insert, or ~ 4 Kbp.
- If you did not get a band, you should use 2-3x the usual recommended DNA amount in your mutant transformation.
- If you did get a band, estimate the approximate amount of DNA in that lane in ng (by comparing to the ladder standards), then the concentration in ng/μL (based on the sample volume that you loaded). Write this information in your notebook.
Part 3: Prepare tubes for liquid O/N cultures
You will make your teaching faculty very happy if you contribute to their preparatory work. Please label 2 large glass test tubes with your team color and sample name in duplicate (X#Z-1, X#Z-2). Normally we would have you combine LB culture broth with ampicillin and aliquot 2.5 mL of this mixture per tube, as these will be used to set up liquid overnight cultures from your two colonies for next time. Due to the long break before your colonies are actually picked, we will do this part for you closer to M1D4.
Part 4: Bacterial transformation
You will transform competent cells called XL1-Blue with your X#Z mutagenesis reactions and plate them on ampicillin-containing Petri dishes. Before next time, two candidate colonies will be chosen from each group's plate. The efficiency of this mutagenesis protocol is reported to be ~80%. We will test two candidates per mutation to cover our bases, so to speak.
- Get an aliquot of competent cells from one of the teaching faculty. Keep these cells on ice at all times, allowing them to thaw slowly (over a few minutes).
- Label three 14 mL polypropylene round-bottom tubes as follows: (-) control, (+) control, X#Z.
- The negative control will receive no DNA, but otherwise go through all the following steps.
- The positive control for you is the pre-tested M124S teaching sample. Normally, a control that comes with the mutagenesis kit we used on Day 2 would be used. Be sure to use a fresh pipet tip when taking from the positive control stock DNA!
- Add 50 μL of competent cells to each tube, followed by 2 μL of the appropriate DNA. Gently swirl (do not vortex) to mix, then incubate on ice for 10 min.
- Bring the tubes over to the 42 °C water bath, and immerse them for exactly 45 seconds according to your digital timer.
- Immediately return the cells to ice for 2 minutes, and take an aliquot of pre-warmed LB medium.
- Add 0.5 mL of warm LB to each sample, then move them to the 37 °C incubator. Ask the teaching faculty to show you how to operate the roller and balance your tubes.
- Allow the cells to recover and begin expressing ampicillin resistance for 30 minutes. At the same time, pre-warm and dry three LB+AMP plates by placing them in the 37°C incubator, media side up with the lids ajar.
- Plate 250 μL of each transformation mix on LB+AMP plates. After dipping the glass spreader in the ethanol jar, you should pass it through the flame of the alcohol burner just long enough to ignite the ethanol. After letting the ethanol burn off, the spreader may still be very hot, and it is advisable to tap it gently on a portion of the agar plate without cells in order to equilibrate it with the agar (if it sizzles, it's way too hot). Once the plates are ready, wrap them together with one piece of colored tape and incubate them in the 37°C incubator overnight. One of the teaching faculty will remove them from the incubator and set up liquid cultures for you to use next time.
Part 5: Practice analysis (optional - but a good idea!)
Ultimately, you will analyze your calcium titration assay data in two steps. First, you will get a rough feel for how your mutants changed (or didn't) compared to wild-type IPC by plotting the two replicate values and their average, in both raw and processed form. Second, you will take the average processed values and plug them into some MATLAB code that will more precisely tell you the affinity and cooperativity of each protein with respect to calcium. The first step is what you will practice today. That way, you can readily plug your own data, acquired on Day 7, into the Excel sheet that you make today and quickly move on to the MATLAB analysis. I will hold office hours to assist with the second step, most likely during the Patriot's day holiday week.
- Open an Excel file for your data analysis. Begin by making a column of the free calcium concentrations present in your twelve test solutions. Assuming a 1:1 dilution of protein with calcium, the concentrations are: 1 nM, 10 nM, 50 nM, 100 nM, 200 nM, 300 nM, 400 nM, 600 nM, 800 nM, 1 μM, 2.5 μM, 10 μM. Be sure to convert all concentrations to the same units.
- Now open the text file containing the practice raw data as a tab-delimited file in Excel; you can download the file from here (TXT download). Convert the row-wise data to column-wise data (using Paste Special → Transpose), and transfer each column to your analysis file. Add column headers to indicate which protein is which, and analyze each replicate separately for now. Also include a column of your control samples that did not contain protein.
- The first two rows are wild-type IPC, the next two T79P, then M124S, and the seventh row is blank (calcium solution).
- Begin by calculating the average of your blank samples, and bold this number for easy reference. It is the background fluorescence present in the calcium solutions and should be quite low. If necessary, subtract this background value from each of your raw data values. It may help to have a 6-column series called “RAW”, and another called “SUBTRACTED.”
- Next you should normalize your data. The maximum and minimum fluorescence values for a given titration series should be defined as 100% and 0% fluorescence, respectively, and every other fluorescence value should be expressed as a percentage in between. Think about how to mathematically express these conditions.
- First calculate the percent fluorescence for both replicates. Then make a new column and calculate the average percentage as well. Alternatively, average your data first, and then normalize the average data.
- If one data point seems really off from the other replicate and from the expected trend, you might consider it an outlier and delete it, especially if you have good reason to believe that there was a reason (error in pipetting, air bubble in that well) for the anomaly. Otherwise, you might be losing valuable information, and/or misleading anyone who tries to interpret your data. (Obviously this point does not apply today, but remember it when you do your analysis for real!)
- For each protein, plot this normalized data versus calcium concentration. Save these plots in case you want to include them in your report.
- You might plot the two replicates as points and their average value as a dashed line (see front page of this module).
- Note down the approximate inflection points of the curves, which should occur at half-saturation: these indicate the approximate values of the apparent [math]\displaystyle{ K_D }[/math] for each sample.
For next time
Revise for S13
1. The major assessment for this module will be a research article describing your protein design work. For this assignment, you will write a draft of the introduction to your report. Recall that the introduction provides a framework for the story you are about to tell, with respect to both background and motivation. You are encouraged to revisit the scientific writing guidelines and brainstorm using the suggested three-section structure before you begin.
- What might the big picture section consist of? Will you focus on engineering sensors in general, the utility of sensing calcium specifically, or ... what?
- What background would the reader like to have as you zoom in?
- When you define your specific study, be sure to share your hypotheses about both your unique mutant and one of the positive control mutants.


2. The pRSET plasmid with inverse pericam insert, or pRSET-IPC, is 4141 basepairs long and its GenBank file is linked here. According to the cutters list that you used on Day 1, restriction site PvuI occurs at ~1655 bp, and again at ~2700 bp into pRSET-IPC. Thus, digesting this parental plasmid with the PvuI enzyme should result in two linear fragments of DNA, with about 1045 and 3095 bp sizes.
A silent mutation can be introduced that results in a new PvuI site at the 341st-342nd residues of inverse pericam (ATT → ATC and TAC → GAC) , or approximately the 1020th basepair of IPC. When IPC is inserted into pRSET, its starting point is ~200bp into the pRSET plasmid. Thus, if the mutated pRSET-IPC plasmid is digested with PvuI, three linear fragments of DNA are the result: 440, 1045, and 2655 bp. To understand these calculations, see also the plasmid maps above. Make sure you can reproduce the numbers in this example before proceeding with your own samples. Plasmid maps can be made using the Enzyme Selector and Graphic Map functions in ApE.
For this assignment, you should plan restriction enzyme digests that allow you to distinguish parental and mutant pRSET-IPC for your X#Z mutation and for your positive control. You are probably best off doing a single enzyme digest for this particular experiment. However, in other kinds of experiments (notably cloning) using two enzymes per digest can give more information. We will discuss these types of digests in a future pre-lab if there is time.
Use the tabulated information on the Day 1 Talk page along with the cutters lists from the Day 1 protocol to determine the cut sites relevant for M124S and T79P. For E67K, the students could not create an enzyme site on the 0- or 1-and-2-cutters lists. Instead, they ended up making a site for a 6-cutter (linked here). The banding pattern should nevertheless be distinguishable for the parent and mutant case, even though not all the bands will be visible.
Please clearly show all your work and reasoning throughout this assignment.
(A) First, use the NEB site to determine the appropriate buffer and temperature for your two reactions.
(B) Next, you should explicitly show how the digest distinguishes between parent and mutant. In other words, what are the expected band sizes for each upon digestion?
(C) Finally, to prepare for Day 4 in lab, calculate the amount of enzyme and buffer needed for your digestion master mixes. Refer to Part 5 (first sub-part) of the Day 4 protocol.
3. Please download the midsemester evaluation form found here (DOC download). Complete the questionnaire and then print it out without including your name to turn in. If there is something you'd like to see done differently for the rest of the course, this is your chance to lobby for that change. Similarly, if there is something you think the class has to keep doing, let us know that too. These will be collected in a separate folder from your FNTs, to preserve anonymity.
4. Extra credit (1 pt): Consider a mutant pRSET-IPC with a newly introduced NotI restriction site that did not occur at all on the parental plasmid (zero cutter). After digestion, you see banding patterns that indicate an uncut plasmid (supercoiled and circular) rather than a linearized piece of DNA for both the parental and the mutant plasmid. Your lab partner tells you that the mutagenesis reaction must not have worked, and that the so-called mutant couldn't possibly contain a NotI site. What’s an equally valid interpretation of this data?
Reagent list
- QuikChange II Site-Directed Mutagenesis Kit from Stratagene
- XL1-Blue supercompetent cells
- LB (Luria-Bertani broth)
- 1% Tryptone
- 0.5% Yeast Extract
- 1% NaCl
- autoclaved for sterility
- Ampicillin: 100 mg/mL, aqueous, sterile-filtered
- LB+AMP plates
- LB with 2% agar and 100 μg/ml Ampicillin
